Designed for identification of new actives against proteins essential for DNA stability
5 760 compounds
For a long time, targeting DNA was considered only in the context of non-specific and cytotoxic agents. Almost all known to date DNA-interacting drugs pose catastrophic actions to the live cells. Several well-known, previously, and still widely used cancer drugs such as Cisplatin, Doxorubicin, and ethidium bromide are non-selective DNA binders or intercalating compounds. Within the last several years protein-DNA interaction interfaces gained much attention among drug hunters. This area is under investigation and requires intensive research to bring new molecules to the clinic. To meet the growing need for new specific DNA binders, we have created a versatile library combining different approaches and tools.
The library has been plated for most convenient and quick access. Using our DNA focused library you receive multiple benefits, allowing you to save on time and costs in hit expansion and optimization:
- Analogs and hit samples resupply from dry stock of over 4.7M compounds
- Straightforward & low-cost synthesis of follow-up libraries through our REAL Database technology
- Medicinal chemistry support enhanced with on-site broad ADME/T panel
Typical Formats
DNA Library is available for supply in various pre-plated formats, including the following most popular ones:
Catalog No.
DNA-5760-0-Z-10
Compounds
5 760
5 plates
Amount
≤ 300 nL of 10 mM of DMSO solutions
Plates and formats
1536-well Echo LDV microplates, first and last four columns empty, 1280 compounds per plate
Price
Catalog No.
DNA-5760-10-Y-10
Compounds
5 760
18 plates
Amount
≤ 10 µL of 10 mM DMSO solutions
Plates and formats
384-well, Echo Qualified LDV microplates #001-12782 (LP-0200), first and last two columns empty, 320 compounds per plate
Price
Catalog No.
DNA-5760-50-Y-10
Compounds
5 760
18 plates
Amount
50 μL of 10 mM DMSO solutions
Plates and formats
384-well, Greiner Bio-One plates #781280, 1,2 and 23,24 columns empty, 320 compounds per plate
Price
Catalog No.
Library & follow-up package
Plates and formats
DNA-5760-10-Y-10 screening library 5 760 cmpds, hit resupply, analogs from 4.7M+ stock and synthesis from REAL Space
Price
*We will be happy to provide our library in any other most convenient for your project format. Please select among the following our standard microplates: Greiner Bio-One 781270, 784201, 781280, 651201 or Echo Qualified 001-12782 (LP-0200), 001-14555 (PP-0200), 001-6969 (LP-0400), C52621 or send your preferred labware. Compounds pooling can be provided upon request.
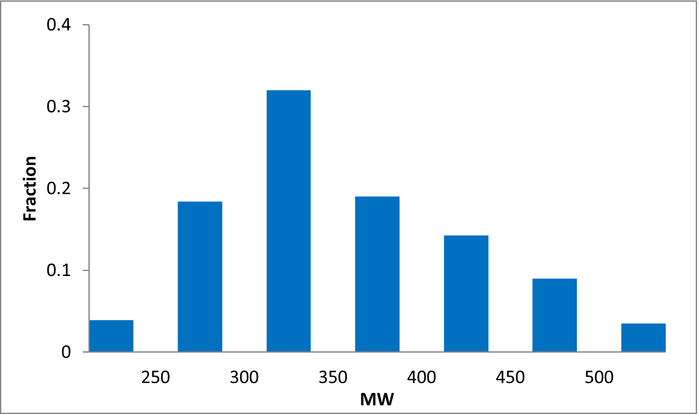
Library design
Different DNA structures have been carefully investigated and analyzed: (1) G-quadruplex structures; G-quadruplexes are noncanonical DNA structures that consist of stacked G-tetrads formed by guanine-rich sequences. These structures are stabilized by monovalent cations such as K+ and Na+. G-quadruplexes are commonly found in the promoter regions of oncogenes like MYC and play an important role in regulating gene expression. Understanding the binding of ligands to G-quadruplexes provides insights into potential therapeutic strategies for targeting oncogenes and modulating gene expression. (2) The double helix DNA structure is a fundamental arrangement where two strands of DNA wind around each other. The integrity of DNA double-strands is crucial for the stability and functioning of cells and organisms. DNA double-strand breaks are particularly critical events that can lead to various consequences. They are associated with the development of human syndromes, neurodegenerative diseases, immunodeficiency, and cancer.
To search for potential DNA ligands all available in PDB and in PDBe liganded DNA structures were analyzed. Then the following entries were selected for in silico screening: 5W77, 6JJ0, 7KBW, 5T4W, 6IJW, 7KWK, 5Z80, 6CCW, 6SX3, and 7EL7. The two main approaches, binding types, were used for search of potential ligands:
- Intercalation: ligands insert itself between the stacked DNA bases pairs.
- Binding insight the DNA grove: ligands binds to the DNA groove without significantly disrupting of the DNA structure.
The molecular docking simulations were conducted using the following queries: pharmacophore of H-bond acceptors and H-bond donors, aromatic group, stearic and volume exclusion features. Each pharmacophore model includes both common and specific features, making the binding points specific to each DNA target structure and its corresponding native ligand. Several examples are provided below.
Examples of molecular docking simulation to histone lysine methyltransferases
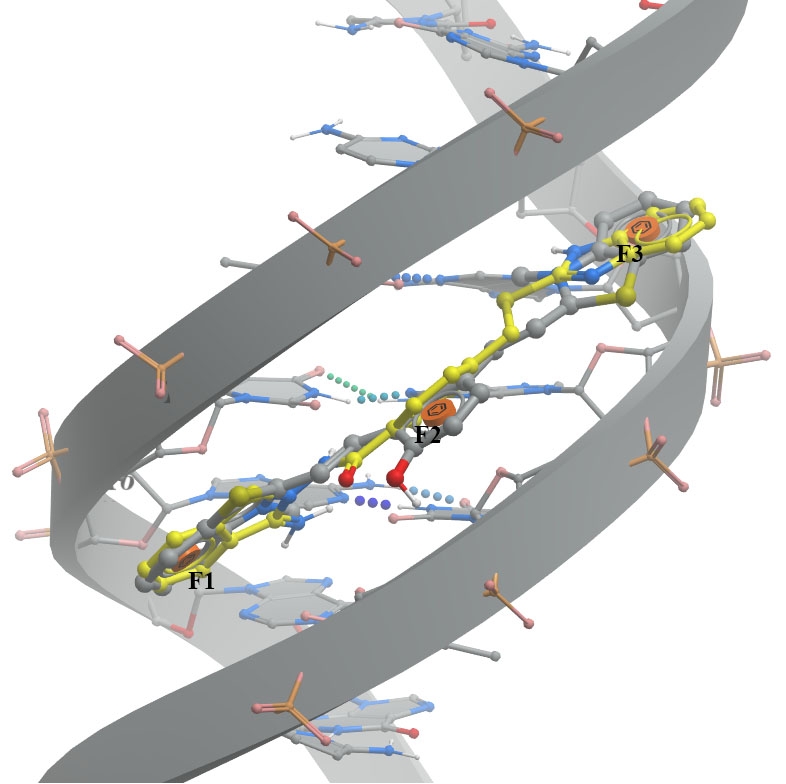
A. The minor groove (PDB ID 6IJW - left image) involves binding between QCy-DT DNA ligand and 6IJW, ligand should match with three aromatic features.
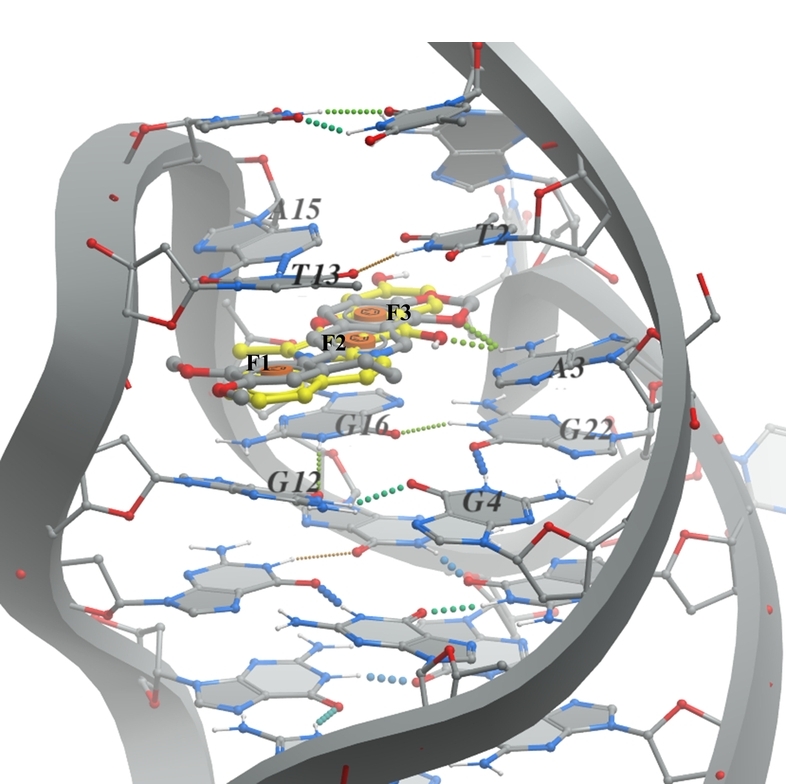
B. G-quadruplex (PDB ID 6CCW - right image) the pharmacophore model based on the presence of epiberberine intercalated within the G-quadruplex. The ligand should consist of three aromatic parts (F1, F2, and F3) that can stack between the DNA nucleobases G12, G16, A15, and T13.
DNA and its native ligand are colored in grey, docked ligands are shown in yellow.
A unique library of most diverse drugs
1 123 compounds
Drug repurposing is a promising field in drug discovery that identifies new therapeutic opportunities for existing drugs (e.g. Metformin and Raloxifene). High-content screens, new biomarkers, and noninvasive imaging techniques have created new capabilities for pursuing novel indications for approved compounds. Hits from our FDA Approved Drug Collection will provide a significant head start in any drug optimization program.
- Carefully collected and distilled collection of 1 123 FDA approved drugs
- Supported with full bioactivity annotation, pathway indications and all related chemical structure information (structure, CAS, smiles, molecular parameters)
- NMR and HPLC validated to ensure the highest purity
- Supplied in 96, 384 or 1536-well plates, ready for immediate shipping
- Minimal preparation – just peel, dilute & transfer to assay plate
Download SD file
Therapeutic areas
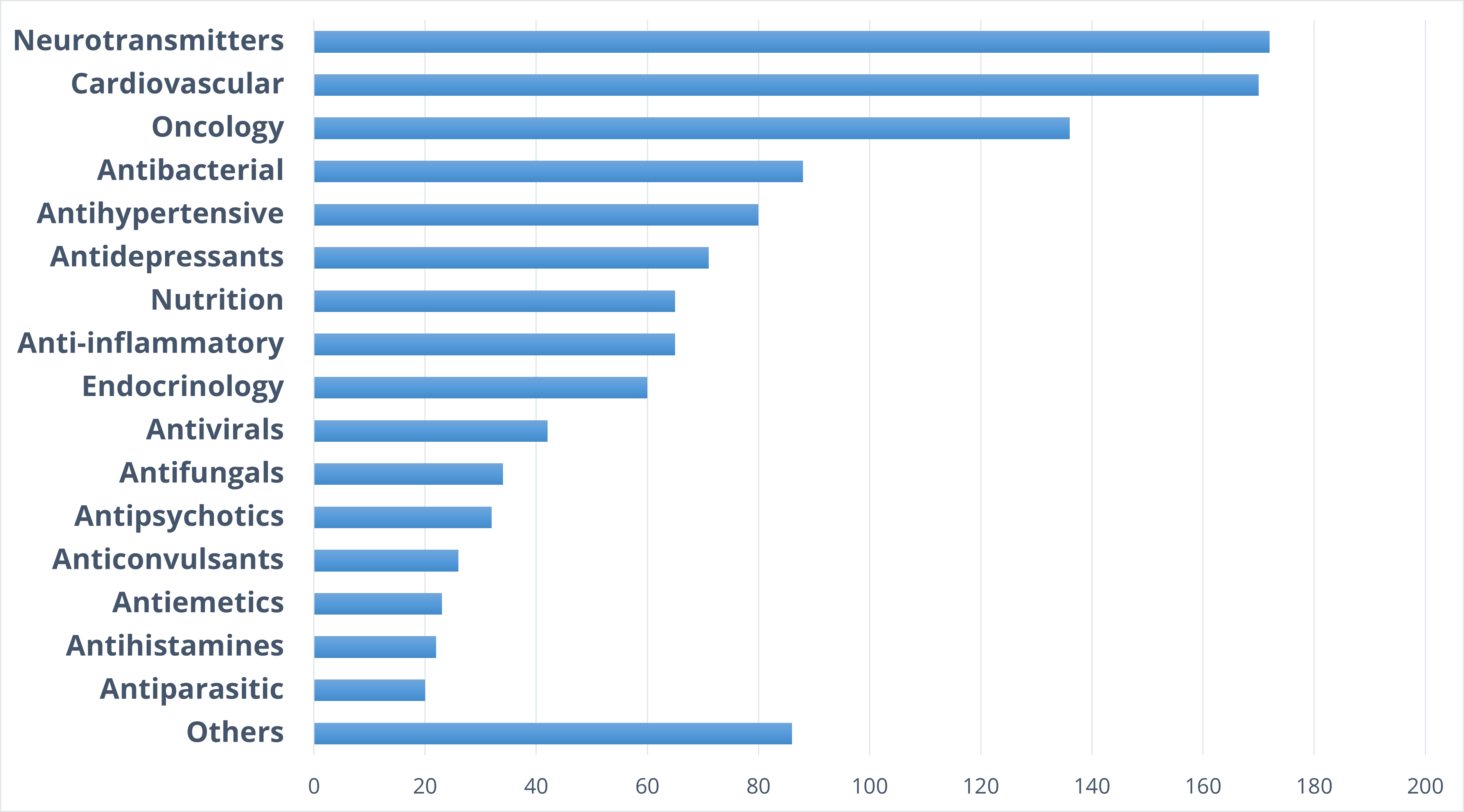
Target distribution
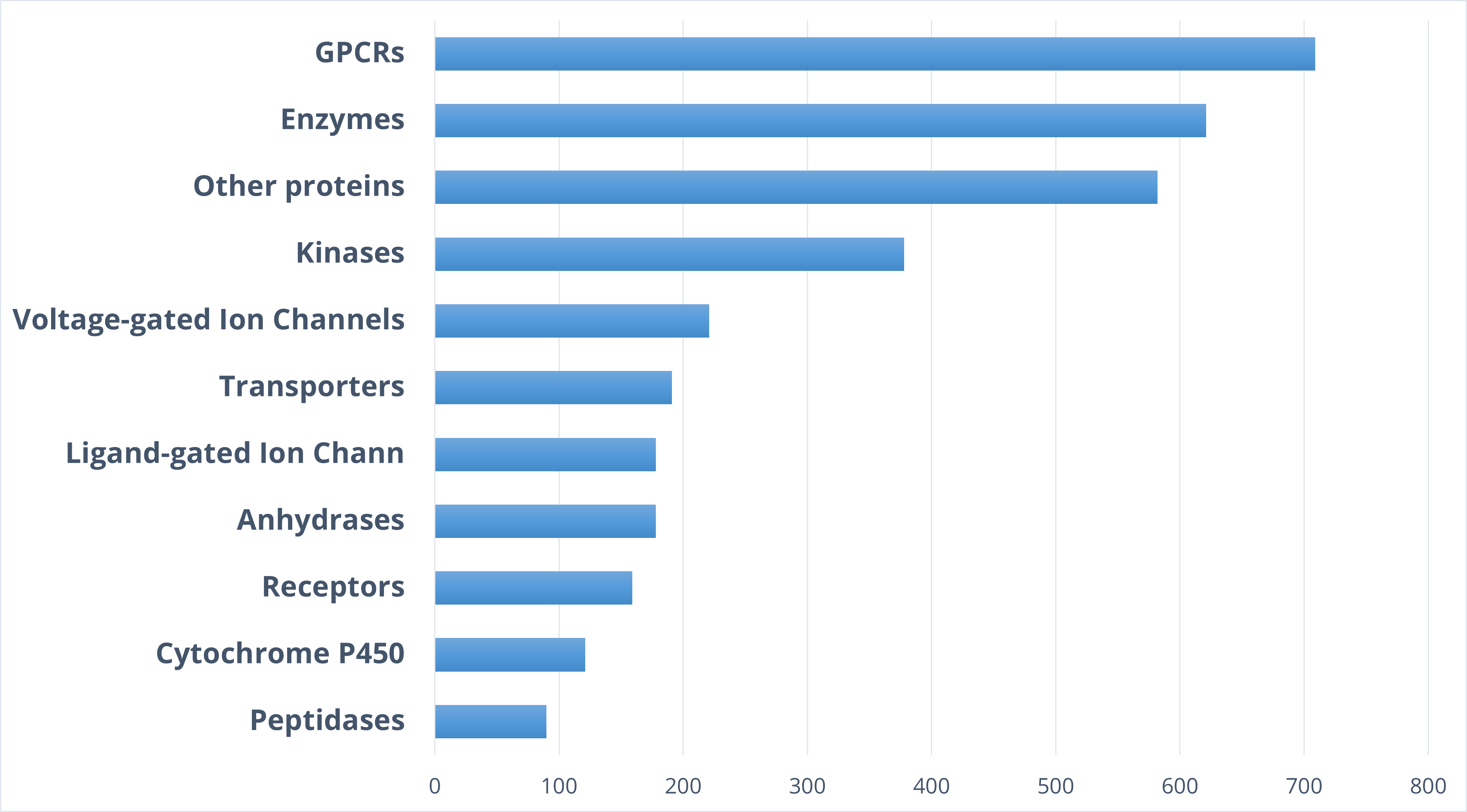
Typical Formats
Catalog No.
FAD-1123-0-Y-2
Compounds
1 123
Amount
≤ 150 nL of 2 mM DMSO solution
Plates and formats
384-well polypropylene plates, first and last two columns empty 320 compounds per plate
Request a quote
Catalog No.
FAD-1123-10-Y-10
Compounds
1 123
Amount
10 µL of 10 mM DMSO solution
Plates and formats
384-well echo-compatible plates: first and last columns empty LP200
Request a quote
Catalog No.
FAD-1123-50-Y/X-10
Compounds
1 123
Amount
50 µL of 10 mM DMSO solution
Plates and formats
384-well or 96-well plates: first and last columns empty
Request a quote
The comprehensive Drug Collection
299 compounds
To date no commercially available compound collection covered all drug classes, resulting in complications in combinatorial screening attempts. We collaborated with a group of researchers at CeMM and collected a condensed screening set of 299 small molecules representing the entire target and chemical space of all FDA-approved drugs. The CeMM Library of Unique Drugs (CLOUD) covers prodrugs and active forms at pharmacologically relevant concentrations and is ideally suited for combinatorial studies. Recent publication [1] provides evidences of successful drug repurposing by pairwise compound screening from CLOUD collection in cancer cell viability assay. The findings highlight that refined chemical library CLOUD is a powerful tool for drug repurposing and evaluation of new substances suitable for all high-content and high-throughput assays.
CLOUD is now available for immediate supply in the following most popular pre-plated formats
Typical Formats
Catalog No.
CLOUD-Y-0
Compounds
299
Amount
≤ 1 µL of 10 mM of DMSO solutions
Plates and formats
384-well echo-qualified assay ready microplates, Greiner/Labcyte
Price
Catalog No.
CLOUD-Y/X-10
Compounds
299
Amount
10 µL of 10 mM of DMSO solutions
Plates and formats
384-well or 96-well plates, echo-compatible
Price
Catalog No.
CLOUD-Y/X-50
Compounds
299
Amount
50 µL of 10 mM of DMSO solutions
Plates and formats
384-well or 96-well plates, echo-compatible
Price
Download SD file
Key features
- Covers all therapeutically significant space of approved drugs.
- Contains prodrugs and active metabolites at pharmacologically relevant concentrations.
- Ideally applicable for combinatorial assays to study new drug combinations.
- Most structurally diverse drug library among commercially available collections.
- Immediately accessible in convenient pre-plated formats with carefully prepared detailed documentation. The library can be also made in any customized ready-to-screen formats.
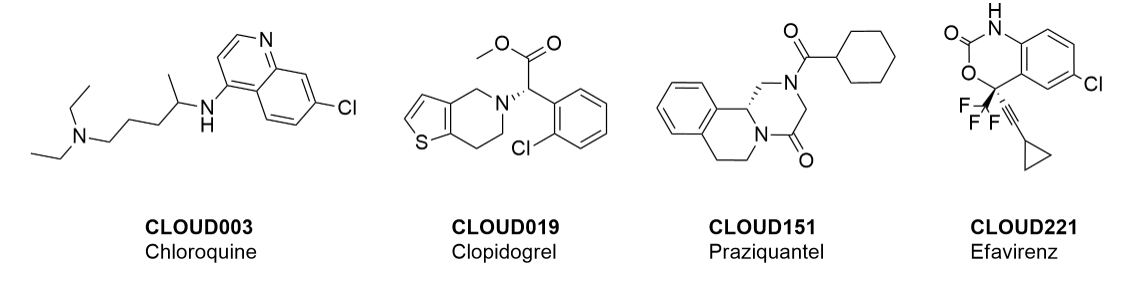
Examples of drugs in the library
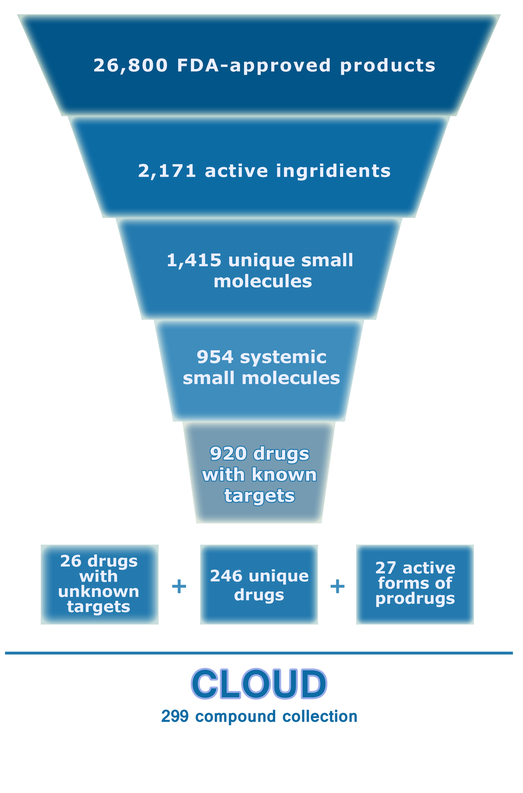
Selection and library evolution scheme

[1] Nature Chemical Biology 13, 771–778 (2017).
Fragments of high MedChem tractability
1 920 compounds
Fragment screening usually provides high numbers of hits and is one of the most affordable approaches to start drug discovery. However, subsequent steps involving hit-to-lead optimization may be challenging and require significant resources. It’s even more painful when efforts result in false positives.
We have designed a new fragment library specifically addressing the quality of decision making when evaluating hits. Our High Fidelity Fragment Library features high medchem tractability enabling researchers to grow interesting hits with confidence.
High medchem tractability of this set was achieved through structure review and selection by FBDD experts at Takeda and Carmot Therapeutics. In particular, we would like to thank Dr. Derek Cole, Dr. Dan Erlanson, Dr. David Lawson and Dr. Xiaolun Wang for their involvement in the design of our High Fidelity Fragment Library.
High quality: All compounds selected for this library passed turbidity tests to assure high solubility in water at 1 mM; all aggregators were filtered out. In addition, the fragments in this set were screened by surface plasmon resonance (SPR) to remove any false positive fragments. SPR screening was kindly provided by Dr. Delphine Collin at HarkerBio.
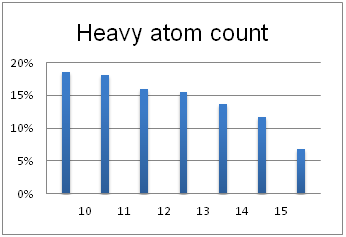
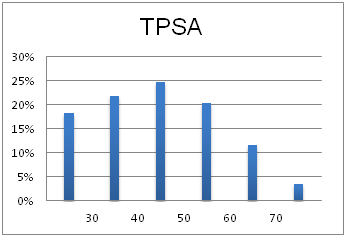
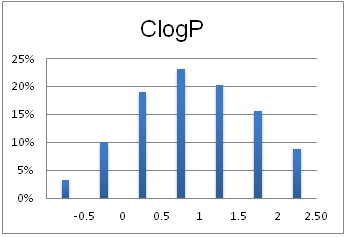
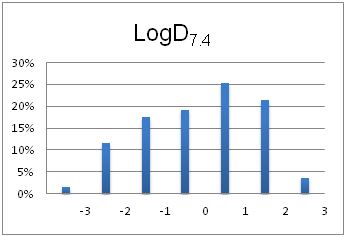
Optimal molecular properties: Fragments in this set all have 9–16 heavy atoms, are moderately complex and have suitable physiochemical and shape profiles.
Our High Fidelity Fragment Library is available for prompt delivery in various formats. The most popular library options are listed below:
Typical Formats
Catalog No.
HFF-1920-Y-10
Compounds
1 920
6 plates
Amount
10 µL of 100 mM DMSO stock solutions
Plates and formats
384-well microplates, Echo qualified Labcyte LP0200
Price
Catalog No.
HFF-1920-Y-25
Compounds
1 920
6 plates
Amount
25 µL of 100 mM DMSO stock solutions
Plates and formats
384-well microplates, Echo qualified Labcyte PP0200
Price
Catalog No.
HFF-1920-X-50
Compounds
1 920
24 plates
Amount
50 µL of 100 mM DMSO stock solutions
Plates and formats
96-well plates, Greiner, first and last columns empty
Price
Download SD file
Library design
A complex iterative approach has been applied in the design of this library. Fragment selection was made from over 260k Ro3-compliant compounds in Enamine’s stock collection. All industry recommended medchem filters, including PAINS, were applied. Diversity selection was performed using clustering algorithm and manual review of identified clusters to select the most representative structure within each cluster.
All compounds were tested for solubility in water and aggregation by laser nephelometry to remove compounds with solubility issues. The resulting set of fragments was then tested in clean SPR screens to exclude SPR “sticky” compounds.
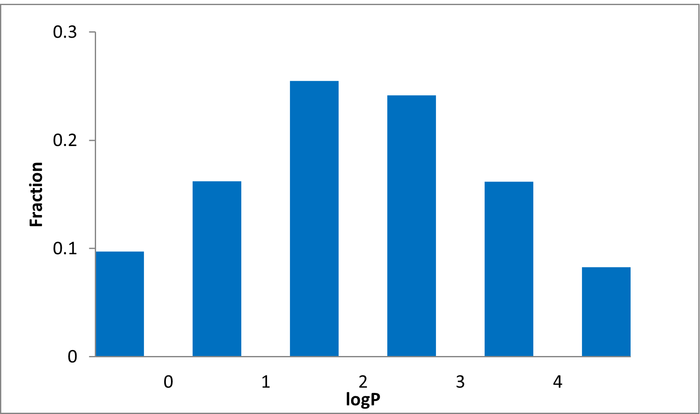
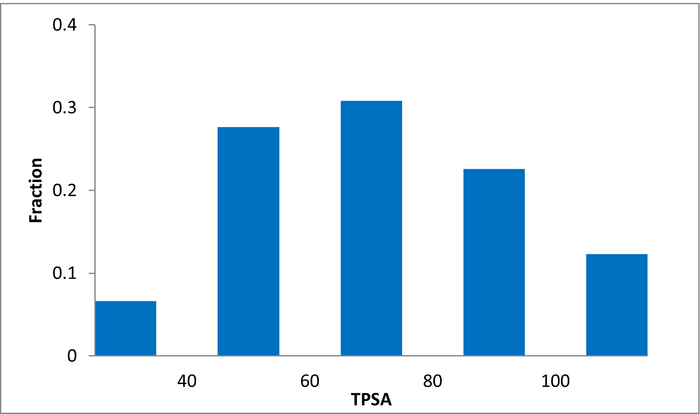
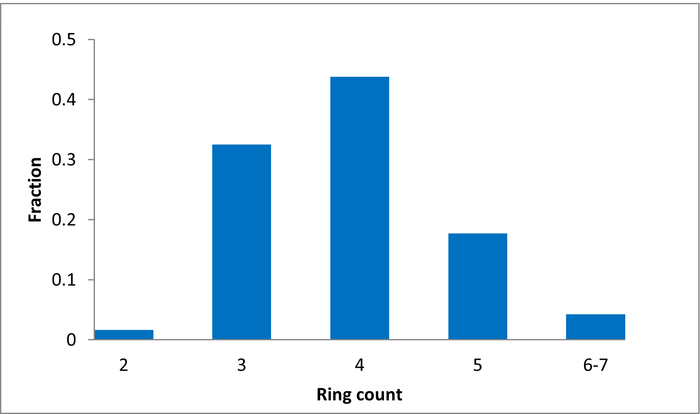
The selection process and resulting molecular parameters of our High Fidelity Fragment Set are described in the following scheme:
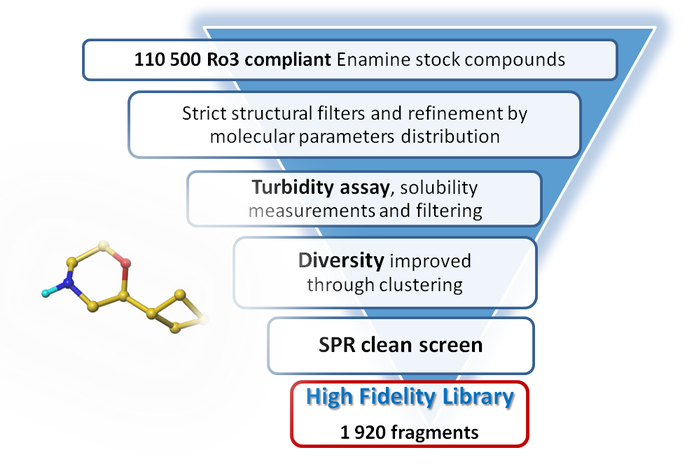
Designed for efficient hit finding against a number of immune disorders, including RA
1 280 compounds
The library was designed to be a universal tool to search potential JAK-STAT pathway modulators. Protein structure-based analysis and scaffold-hopping approach were used to create an optimal in silico screening models. Additionally, ligand-based approach has been applied to enrich the library with topological analogs and similar compounds to reported actives.
Resulted library is as a generic starting point for ligand search, regardless of particular SH2 domain of interest, with high probability of initial hit discovery. The library was validated with in vitro screening against SRC SH2 domain resulting in 3.5 % of identified hits.
Library design
All available structures of SH2 domain binding sites were analyzed and clustered based on their spatial molecular shape. After the alignment analysis and clustering structures with most different conformations were selected for virtual screening. Enamine MedChem filtered in-stock subset (~1 M compounds) with additional selection of compounds carrying peptidomimetic motifs was used for molecular docking calculations.
Multiple sequence alignment was applied to 120 SH2 domains contained within 110 proteins. Over 200 protein structures were extracted from Protein Data Bank (PDB): 66 NMR-based structures and 153 derived from X-ray crystallography experiments. As variation of the binding site conformation may significantly influence its binding properties, all files were split into individual structures, resulting in total structure count of 1633.
Key pharmacophore interaction points: pTyr binding pocket, carbonyl O in the binding site center, hydrophobic sub-pocket.
Solvent accessible molecular surface within 12 Å from the binding site center was used for calculation of shape-based numeric descriptors. 3D structures were clustered based on 3D shape similarity. 8 spatially diverse structures were selected for docking.
Table 1. Summary of protein 3D structures clustering results and structures selected for docking.
1
Centroid structure (PDB id, chain, NMR model)
1o49, chain A
Centroid structure: organism, gene, domain
Homo Sapiens SRC SH2
Structures in cluster
223
2
Centroid structure (PDB id, chain, NMR model)
2fci, chain A, model 6
Centroid structure: organism, gene, domain
Bos Taurus PLCG1 SH2-2
Structures in cluster
48
3
Centroid structure (PDB id, chain, NMR model)
2ge9, chain A, model 15
Centroid structure: organism, gene, domain
Homo Sapiens BTK SH2
Structures in cluster
72
4
Centroid structure (PDB id, chain, NMR model)
3in7, chain A
Centroid structure: organism, gene, domain
Homo Sapiens GRB2 SH2
Structures in cluster
183
5
Centroid structure (PDB id, chain, NMR model)
2jyq, chain A, model 9
Centroid structure: organism, gene, domain
Homo Sapiens GRB2 SH2
Structures in cluster
114
6
Centroid structure (PDB id, chain, NMR model)
2k7a, chain B, model 5
Centroid structure: organism, gene, domain
Mus Musculus ITK SH2
Structures in cluster
218
7
Centroid structure (PDB id, chain, NMR model)
2kk6, chain A, model 14
Centroid structure: organism, gene, domain
Homo Sapiens FER SH2
Structures in cluster
149
8
Centroid structure (PDB id, chain, NMR model)
1uus, chain A
Centroid structure: organism, gene, domain
Dictyostelium Discoideum DSTA SH2
Structures in cluster
129
Virtual screening against Stat3beta:
- Collaborator: Gyeong Baeg, NYMC
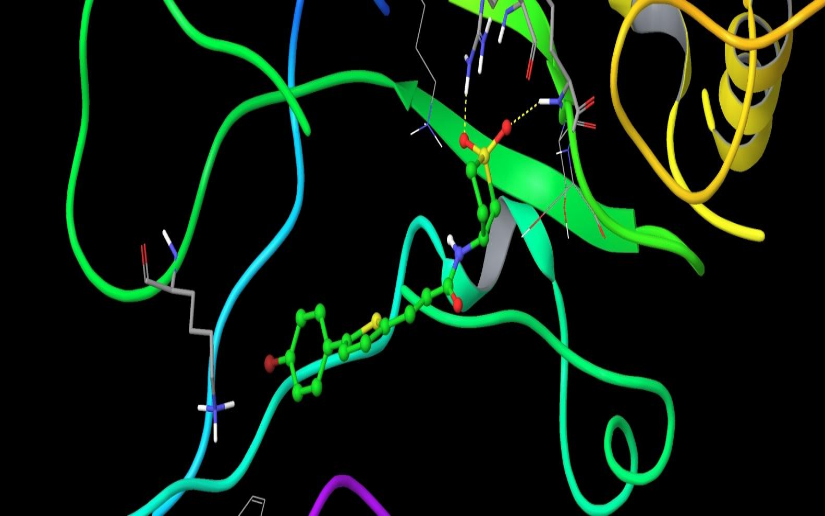
Chemical compounds are significantly smaller in size than the natural peptide interacting with this target. Therefore, when creating the library, we tried to fill all potential sub-pockets available near the phosphotyrosine binding site. The entire available surface of the protein was conditionally broken down into 5 models and proceeding from this was carried out post-docking analysis. As an example, the first two models:
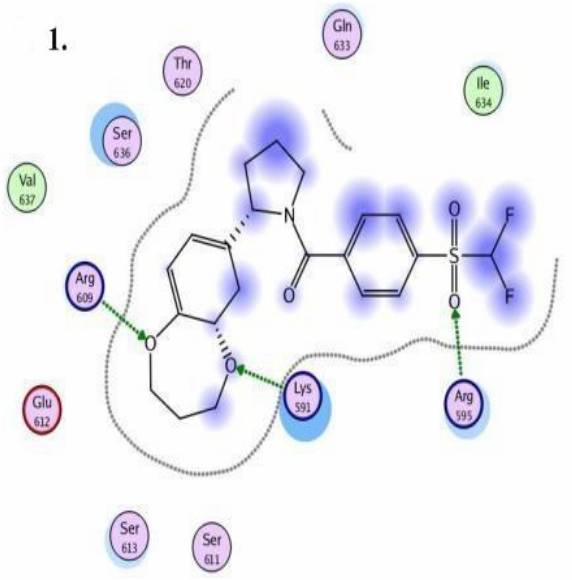
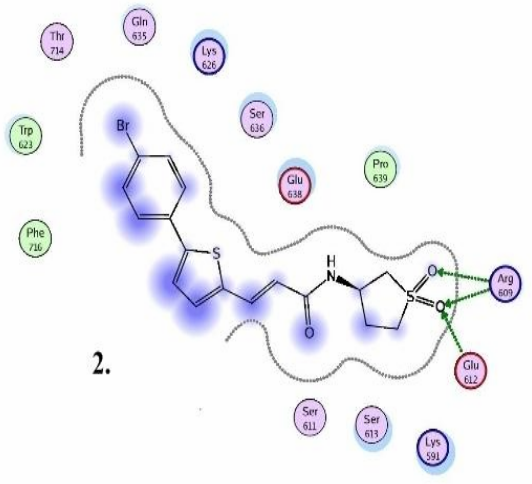
STAT3 models: Schematic representation of ligand binding pocket