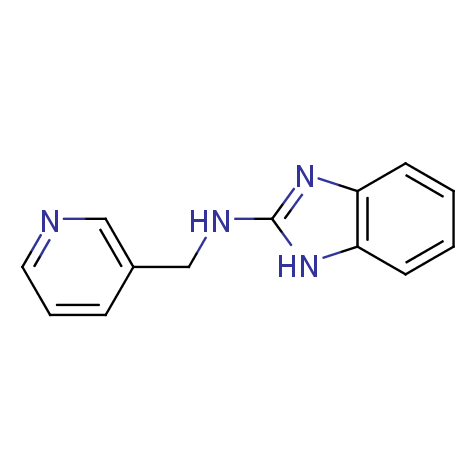IDO focused library designed by a combination of structure- and ligand-based methods
4 800 compounds
Indoleamine 2,3-dioxygenase (IDO) is a heme-containing protein that catalyzes the oxidative cleavage of the C2−C3 double bond of the indole in tryptophan to provide N-formylkynurenine.
Inhibition of this pathway is perspective and very attractive for the treatment of many neurological disorders, such as Alzheimer’s disease, Parkinson’s disease and cerebral ischemia.
In addition overexpression of IDO1 has been reported in many tumor cells that lead to consideration of IDO1 as one of the key factors that contributes to cancer immunosuppression in tumor microenvironment.
Download SD files
Library code: IDO-4800
Version: 13 August 2024
4 800 compounds at 10 mM in DMSO
Library design
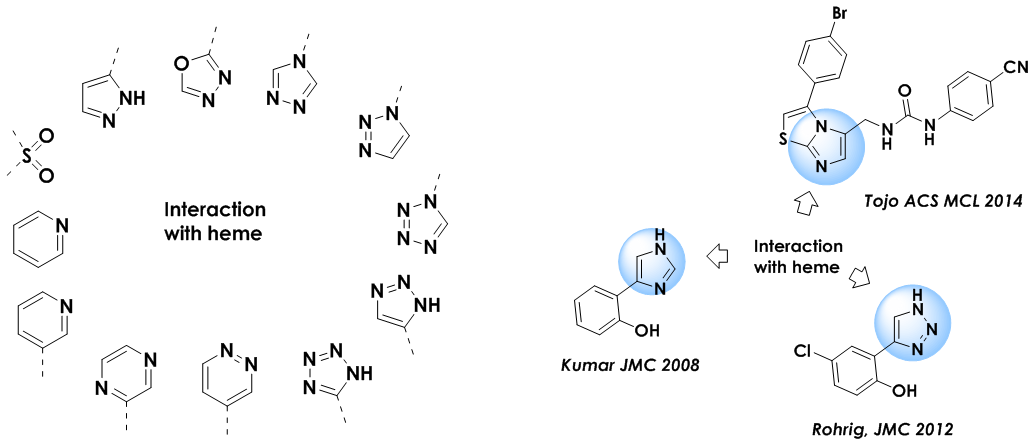
According to the literature data, the key interaction of IDO inhibitors with the biological target is coordination to Fe ion of the heme in the active site. It is this interaction which prevents participation of the Fe ion in indole amine oxidation.
Significant number of the known IDO inhibitors is hydrophilic compounds that have relatively low molecular weight. Therefore, strict PhysChem parameter filters (lead-likeness, Veber rules) were applied for initial compound set selection. We have developed a holistic approach to the library design that includes ligand-/receptor-based virtual screening and enrichment with heme-binding fragments along with new chemotypes selection.
- Ligand-based approach: Compounds with similar scaffolds, chemotypes and key moieties were picked out from our stock collection.
- Docking: Virtual screening was carried out with heme coordination constraint and other important structural features that are specific for vast majority of potent ligands. The ability of ligands to fill the ambient sub-pockets was set as preferable.
- Selection by Substructure Search: Critical search of molecules bearing fragments reported to form coordination bonds with heme. Compounds with unique chemotypes and appropriate shapes were included to the library.
- Molecular properties: 100% Ro5 compliant, and 99.5 % lead-like.
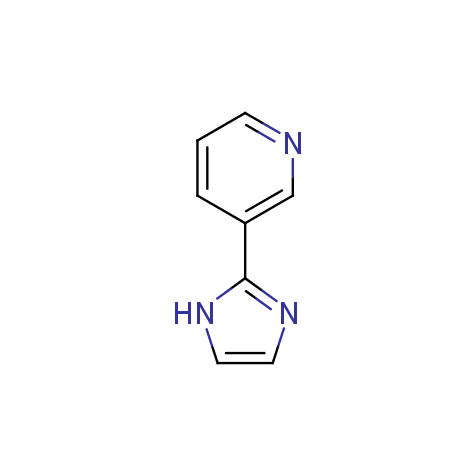
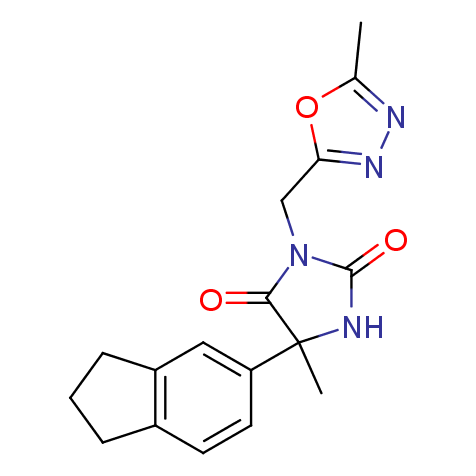
Examples of the molecules in the library
Chelating fragments included in the library and examples of utilized reference compounds