A sub-library of Enamine’s GPCR Library designed for discovery of novel lipid GPCR ligands
6 400 compounds
Enamine’s Lipid GPCR Library has been designed in collaboration with NQuiX. Over 6 400 compounds have been selected from off-the-shelf collection or specifically synthesized for this project from Enamine REAL screening Collection. They were designed to target the family of eight endothelial differentiation gene (EDG) receptors (S1P1–5 and LPA1–3). The compounds may also be appropriate for screening at GPR3, GPR6 and GPR12 orphan receptors given some TM bundle binding site similarity and/or GPR23, GPR92 and P2Y5 given their more recent classification as additional, albeit distinct, LPA receptors.
Typical Formats
Lipid GPCR Library is available for supply in various pre-plated formats, including the following most popular ones:
Catalog No.
LGR-6-0-Z-10
Compounds
6 400
5 plates
Amount
≤ 300 nL of 10 mM of DMSO solutions
Plates and formats
1536-well Echo LDV microplates, first and last four columns empty, 1280 compounds per plate
Price
Catalog No.
LGR-6-10-Y-10
Compounds
6 400
20 plates
Amount
≤ 10 µL of 10 mM DMSO solutions
Plates and formats
384-well, Echo Qualified LDV microplates #001-12782 (LP-0200), first and last two columns empty, 320 compounds per plate
Price
Catalog No.
LGR-6-50-Y-10
Compounds
6 400
20 plates
Amount
50 μL of 10 mM DMSO solutions
Plates and formats
384-well, Greiner Bio-One plates #781280, first and last two columns empty, 320 compounds per plate
Price
Catalog No.
Library & follow-up package
Plates and formats
LGR-6-10-Y-10 screening library 6 400 cmpds, hit resupply, analogs from 4.7M+ stock and synthesis from REAL Space
Price
*We will be happy to provide our library in any other most convenient for your project format. Please select among the following our standard microplates: Greiner Bio-One 781270, 784201, 781280, 651201 or Echo Qualified 001-12782 (LP-0200), 001-14555 (PP-0200), 001-6969 (LP-0400), C52621 or send your preferred labware. Compounds pooling can be provided upon request.
Download SD file
Library design
The compounds have been selected using a combination of ligand-based methods using chemical fingerprints, 2D pharmacophores and 3D shape/feature matches (Figure 1). The training active compound set included 1 393 compounds (870 acidic and 523 non-acidic), collected across 73 papers (Figure 2). They represent a combination of compounds for expanding SAR around known chemotypes and scaffold hops seeking novelty. The current set of compounds covers the non-acidic classes of ligand, with acidic compounds being designed via a different procedure and added to the library separately.
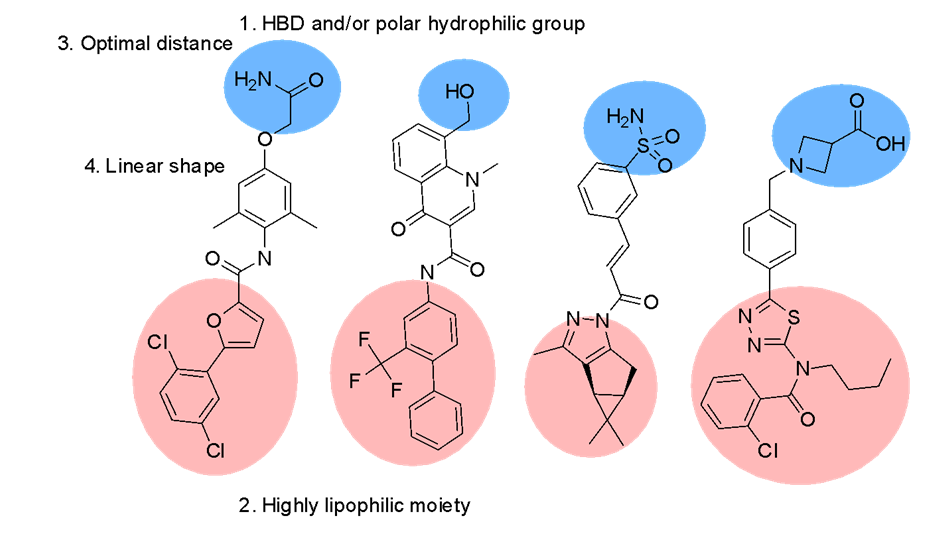
Fig. 1.Dedicated design of the compounds included in the Library (general scheme).
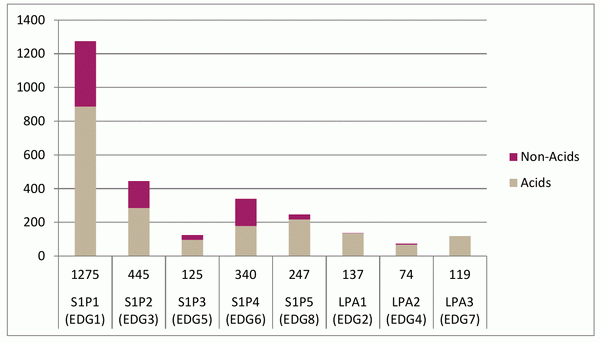
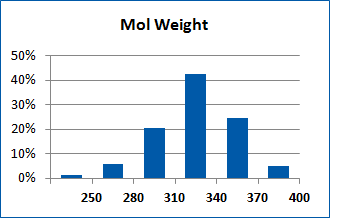
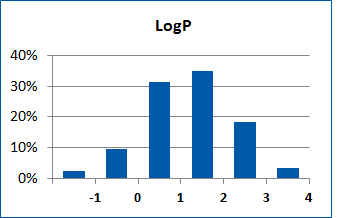
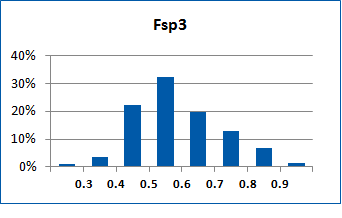
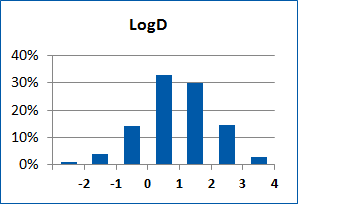
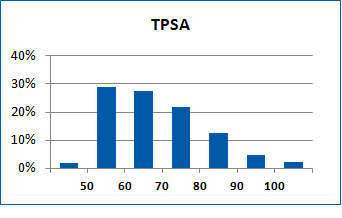
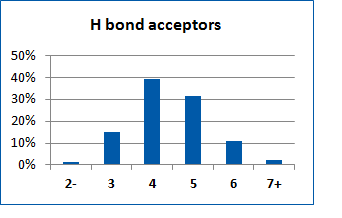
Fig. 2. Distribution of the compounds in the training set.
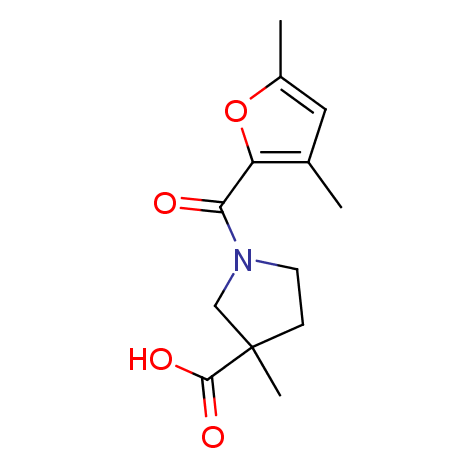
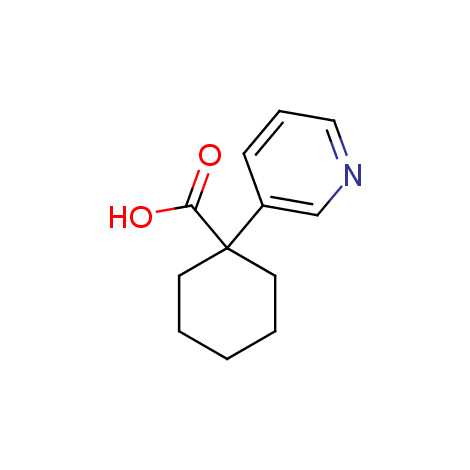
Examples of the molecules
Designed for discovery of novel allosteric ligands
14 160 compounds
Although modulating GPCRs with orthosteric selective drug candidates has numerous success stories, in many cases, allosteric GPCR ligands are more effective for these biological targets. The small molecule allosteric modulation of GPCRs can promote a conformational change in the receptor that often has no noticeable effects, but can have a strong impact on the signaling pathway in the presence of an orthosteric endogenous ligand. On the other hand, GPCRs actuated by intractable stimuli can be modulated allosterically, opening a new opportunity for these currently unassailable targets. Two allosteric GPCRs modulators that have been introduced in the clinic (Cinacalcet and Maraviroc) are excellent evidence of prospects of investigation in this area. Another benefit of these ligands is their relatively low molecular weight and hence higher potential for oral bioavailability, in contrast to most orthosteric ligands. Moreover, allosteric sites are generally less conserved enabling development of actives with greater subtype selectivity than drugs targeting the conserved binding site.
Enamine’s Allosteric GPCR Library is a utile starting point for the drug discovery in the field of allosteric GPCR modulators.
Typical Formats
Allosteric GPCR Library is available for supply in various pre-plated formats, including the following most popular ones:
Catalog No.
AGR-14-0-Z-10
Compounds
14 160
12 plates
Amount
≤ 300 nL of 10 mM of DMSO solutions
Plates and formats
1536-well Echo LDV microplates, first and last four columns empty, 1280 compounds per plate
Price
Catalog No.
AGR-14-10-Y-10
Compounds
14 160
45 plates
Amount
≤ 10 µL of 10 mM DMSO solutions
Plates and formats
384-well, Echo Qualified LDV microplates #001-12782 (LP-0200), first and last two columns empty, 320 compounds per plate
Price
Catalog No.
AGR-14-50-Y-10
Compounds
14 160
45 plates
Amount
50 μL of 10 mM DMSO solutions
Plates and formats
384-well, Greiner Bio-One plates #781280, 1,2 and 23,24 columns empty, 320 compounds per plate
Price
Catalog No.
Library & follow-up package
Plates and formats
AGR-14-10-Y-10 screening library 14 160 cmpds, hit resupply, analogs from 4.7M+ stock and synthesis from REAL Space
Price
*We will be happy to provide our library in any other most convenient for your project format. Please select among the following our standard microplates: Greiner Bio-One 781270, 784201, 781280, 651201 or Echo Qualified 001-12782 (LP-0200), 001-14555 (PP-0200), 001-6969 (LP-0400), C52621 or send your preferred labware. Compounds pooling can be provided upon request.
Download SD file
Library design
Both ligand- and structure-based methods were used in design of the library. Ligand-based approach relied on using 3D pharmacophore search to the reference set of known ligands with high activity values (over 200 compounds from ChEMBL and BindingDB). The main emphasis has been done on allosteric modulators of class B GPCRs that are well established in drug discovery. Two pharmacophore models were created using the reference compounds sets and then validated with the training sets of actives and non-active ligands (Figure 1). Finally, over 6 200 small molecules were selected into Pharmacophore-based allosteric GPCR subset starting from Enamine MedChem set (710 K).
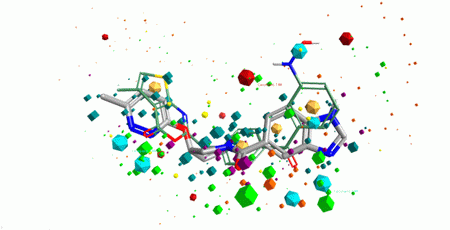
Fig. 1. Pharmacophore superposition of a reference active compound (in green) and a hit compound from the Library (in grey).
Structure-based approach was applied to human Glucagon receptor (GCGR) and human Corticotropin-releasing factor 1 receptor (CRF1R). The corresponding protein structures recorded in 5EE7 and 4K5Y PDB entries were optimized and used for the docking calculation with Glide (Schrödinger software). Compounds with at least 50% score-based efficacy as compared to the co-crystallized ligands were selected as hits and included in the targeted sets: h-GCGR set – 200 compounds; h-CRF1R set: 7 200 compounds (Figures 2 and 3).
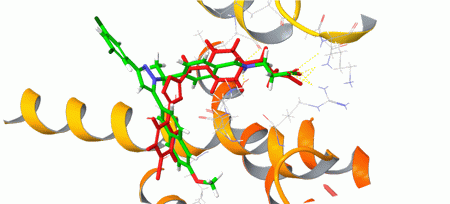
Fig. 2. Binding mode of a hit (in red) obtained from the docking and the co-crystallized ligand (in green) of human Glucagon receptor.
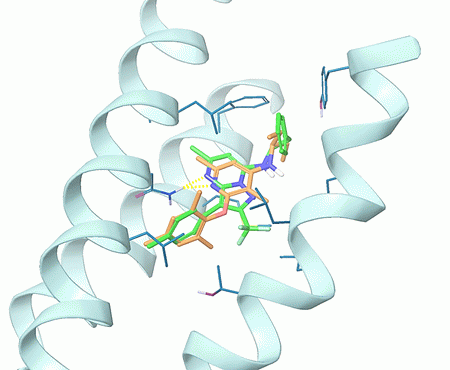
Fig. 3. Superposition of a hit compound obtained after docking of CRF1R allosteric site with Enamine MedChem screening compounds (in green) and the co-crystallized ligand CP-376395 (in orange).
Examples of the molecules
Designed for discovery of novel kinase ATP pocket binders
24 000 compounds
The most straightforward approach to the design of kinase inhibitors relies on targeting ATP pocket. In fact, all compounds of this class approved by FDA (Copanlisib, Neratinib, Brigatinib, and Ribociclib) demonstrate this binding mode. The standard kinase interaction pattern consists of a hydrogen bond acceptor for the hinge region, heteroaromatic core with various substituents, and a second hydrogen acceptor for the conserved Lysine residue. This type of Inhibitors can be considered as ATP mimetics in the sense that they make interactions similar to that of ATP. Hydrogen bonds with Hinge Region formed by the adenosine moiety of ATP are crucial for effective binding (Figure 1, left). Analysis of a broad number of the kinase-inhibitor interactions has shown that similar hydrogen bonding patter is necessary for high inhibitory potency.
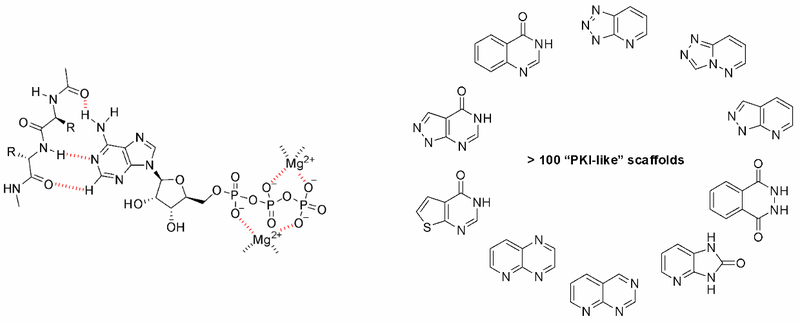
Fig. 1. ATP binding mode with hinge region (left) and examples of core structures that can mimic that mode (right).
Typical Formats
Hinge Binders Library is available for supply in various pre-plated formats, including the following most popular ones:
Catalog No.
HBL-24-0-Z-10
Compounds
24 000
19 plates
Amount
≤ 300 nL of 10 mM of DMSO solutions
Plates and formats
1536-well Echo LDV microplates, first and last four columns empty, 1280 compounds per plate
Price
Catalog No.
HBL-24-10-Y-10
Compounds
24 000
75 plates
Amount
≤ 10 µL of 10 mM DMSO solutions
Plates and formats
384-well, Echo Qualified LDV microplates #001-12782 (LP-0200), first and last two columns empty, 320 compounds per plate
Price
Catalog No.
HBL-24-50-Y-10
Compounds
24 000
75 plates
Amount
50 μL of 10 mM DMSO solutions
Plates and formats
384-well, Greiner Bio-One plates #781280, 1,2 and 23,24 columns empty, 320 compounds per plate
Price
Catalog No.
Library & follow-up package
Plates and formats
HBL-24-10-Y-10 screening library 24 000 cmpds, hit resupply, analogs from 4.7M+ stock and synthesis from REAL Space
Price
*We will be happy to provide our library in any other most convenient for your project format. Please select among the following our standard microplates: Greiner Bio-One 781270, 784201, 781280, 651201 or Echo Qualified 001-12782 (LP-0200), 001-14555 (PP-0200), 001-6969 (LP-0400), C52621 or send your preferred labware. Compounds pooling can be provided upon request.
Download SD file
Library design
Starting from the results of our previous study (Anticancer Agents Med. Chem. 2007, 7, 171–188) and detailed structure analysis of known and most potent kinase inhibitors, we have developed a series of unique structure filters aimed to identify potential inhibitors targeting ATP pocket. Molecular fragments able to form at least two hydrogen bonds with hinge region (Figure 1, right) have been analyzed, and appropriate topological models were developed for searching of directed inhibitors. The screening models were evaluated with a reference set of known kinase inhibitors (over 2 000 molecules with high inhibitory activity). Validated topological models were applied to over 3.2 M Enamine stock collection to produce Kinase Hinge Binders Libarary. The resulting set of compounds was evaluated with main emphasis on novel chemotypes. MedChem filters including PAINS and Ro5 PhysChem restrictions were applied as a prerequisite.
Examples of the molecules
Designed for specific protein targets and sensible onset
4 000 compounds
In response to high demand for new and small carboxylic acid we have created set of carefully selected specific molecules that strictly respond to FBDD requirements. Our carboxylic acid fragments are mainly consist from the compounds derived in the last three years from previously established at Enamine in-house program for design and producing diverse carboxylic acids to enlarge core Building Blocks Collection and diversification of decorating reagents.
Mainly this type of fragments has great potential for hit finding on new and difficult targets, e.g. protein-protein interaction, due to binding affinity to so called “hot-spot” residues. In addition there is probability of high solubility and good membrane permeability of discovered initial hit that often could be a limiting factor for phenotypic screening or in vivo activity of following investigation.
Typical Formats
Catalog No.
CAF-4-10-Y-100
Compounds
4 000
13 plates
Amount
10 µL of 100 mM solutions in DMSO
Plates and formats
384-well plates, Echo qualified Labcyte LP0200; 1, 2 and 23, 24 columns empty; 320 compounds per plate
Price
Catalog No.
CAF-4-50-X-100
Compounds
4 000
50 plates
Amount
50 µL of 100 mM solutions in DMSO
Plates and formats
96-well plates, Greiner Cat. No 650201, 1 & 12 columns empty, 80 compounds per plate
Price
Catalog No.
CAF-4-100-X-10
Compounds
4 000
50 plates
Amount
100 µL of 10 mM solutions in DMSO
Plates and formats
96-well plates, Greiner Cat. No 650201, round (U) bottom, 1 & 12 columns empty, 80 compounds per plate
Price
Download SD file
Key features
- High novelty and core diversity
- Fast follow-up with stock available analogues & synthesis
- Cherry-picking is available, customer preferred delivery format
Examples of the molecules in the library
Designed for discovery of new Ion Channels ligands
36 800 compounds
Ion channels are most widely distributed in organism tissues and play an important role in numerous cell types as large families of related genes with cell-specific expression pattern. This type of therapeutic targets is most important in developing of new highly effective medical treatments in the same time being one of the most difficult tasks in Medicinal Chemistry.
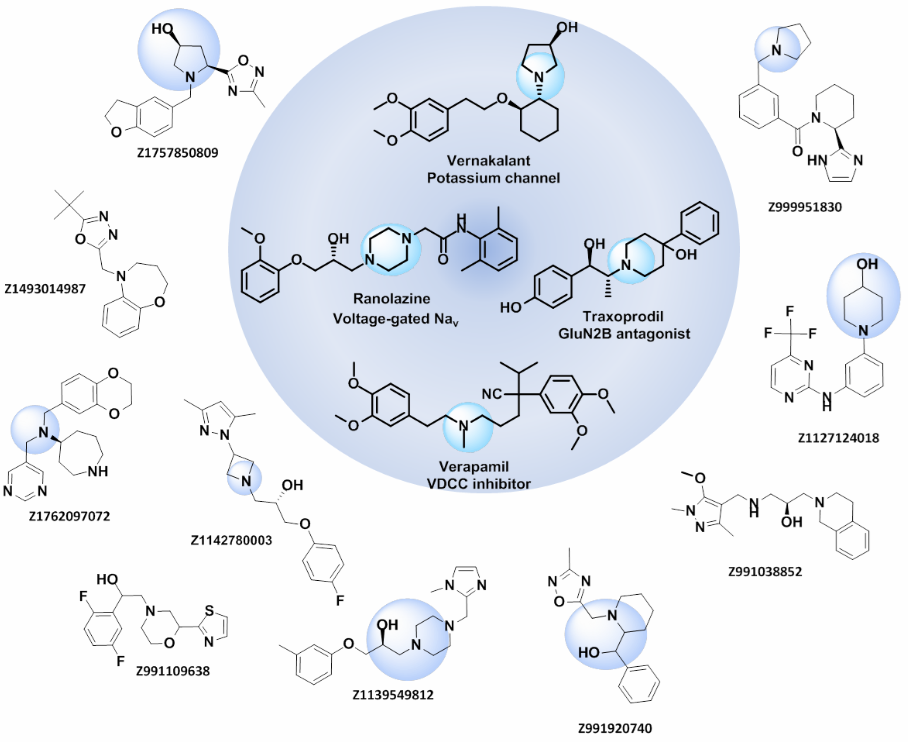
Examples of the molecules
Using our Libraries for hit identification you receive multiple benefits allowing you to save on time and costs in lead generation:
- Resupply from dry powders for hit confirmation, hit expansion from in-stock analogs.
- Straightforward and low-cost synthesis of analogues through our REAL Database technology.
- Medicinal chemistry support enhanced with on-site broad ADME/T panel.
Typical Formats
Ion Channel Library is available for supply in various pre-plated formats, including the following most popular ones:
Catalog No.
ICL-36-0-Z-10
Compounds
36 800
29 plates
Amount
≤ 300 nL of 10 mM of DMSO solutions
Plates and formats
1536-well Echo LDV microplates, first and last two columns empty, 1280 compounds per plate
Price
Catalog No.
ICL-36-10-Y-10
Compounds
36 800
115 plates
Amount
≤ 10 µL of 10 mM DMSO solutions
Plates and formats
384-well, Echo Qualified LDV microplates #001-12782 (LP-0200), first and last two columns empty, 320 compounds per plate
Price
Catalog No.
ICL-36-50-Y-10
Compounds
36 800
115 plates
Amount
50 μL of 10 mM DMSO solutions
Plates and formats
384-well, Greiner Bio-One plates #781280, first and last two columns empty, 320 compounds per plate
Price
Catalog No.
Library & follow-up package
Plates and formats
ICL-36-10-Y-10 screening library 36 800 cmpds, hit resupply, analogs from 4.7M+ stock and synthesis from REAL Space
Price
*We will be happy to provide our library in any other most convenient for your project format. Please select among the following our standard microplates: Greiner Bio-One 781270, 784201, 781280, 651201 or Echo Qualified 001-12782 (LP-0200), 001-14555 (PP-0200), 001-6969 (LP-0400), C52621 or send your preferred labware. Compounds pooling can be provided upon request.
Download SD files
Library design
We applied multipronged approach in rational library design to gain a qualitative set of molecules focused on ion channel targets. A major contribution was made in the lead-oriented synthesis program at Enamine focused on increasing novelty and structural diversity. The project has already yielded 36 800 lead-like compounds built on novel scaffolds featuring saturated rings that have been recognized as potential ion channel blockers. Pharmacophore ligand-based biased analysis was performed on the reference set of over 1 000 highly active ligands (≤10 nM) resulting in three main pharmacophore motifs frequently occurring in reported ligands. Optimized models were used in searching a lead-like part of Enamine Screening Collection and a lead-like part of REAL database. One of the common features of derived pharmacophores was presence of tertiary/secondary amino group. Additional compounds were added to the library after analysis of privileged motives of known ion channel blockers and after morphing of some recently discovered ion channel and TRPV1 modulators. General Medicinal Chemistry overview finalized the library profile: favorable physicochemical parameters and solubility requirements.