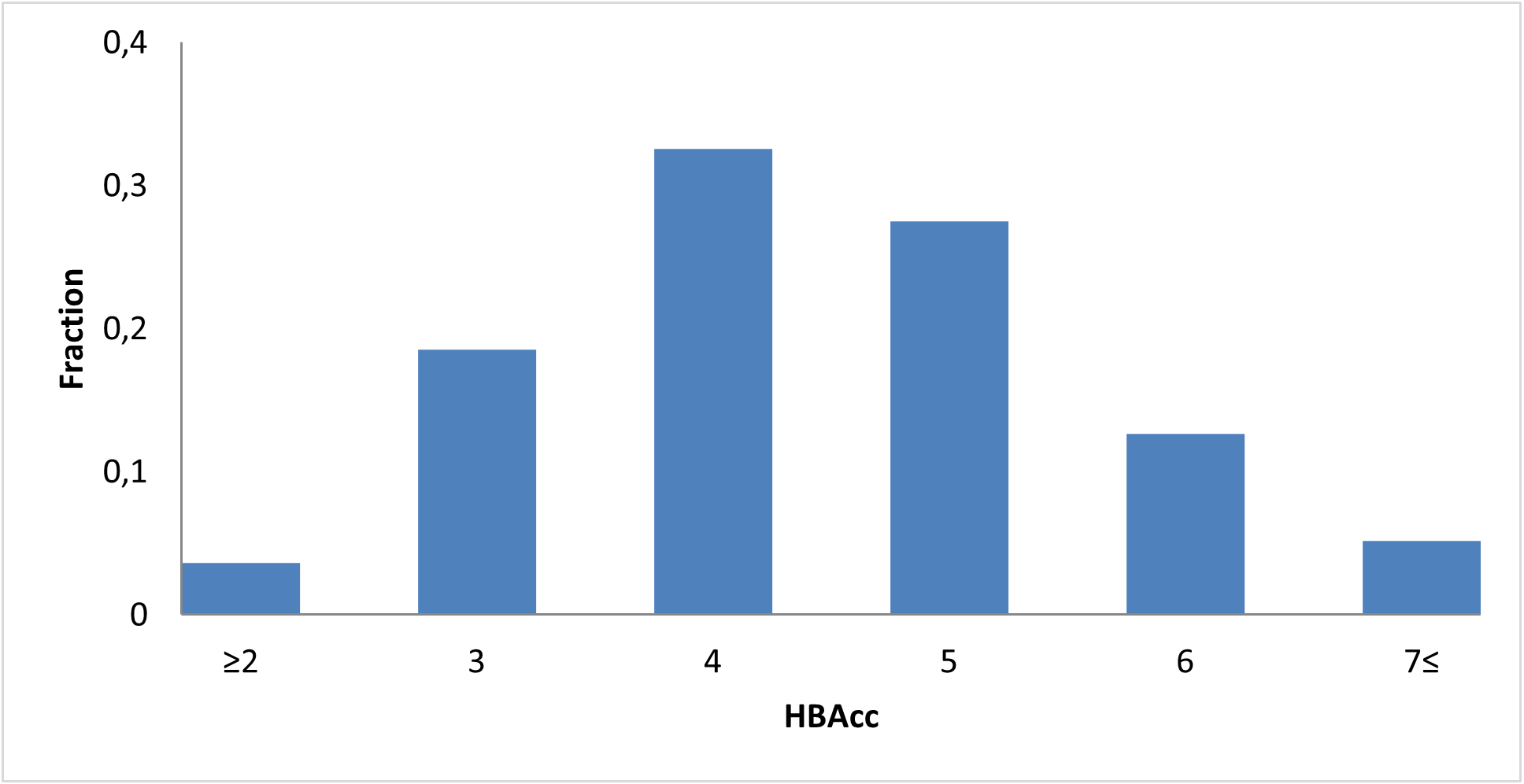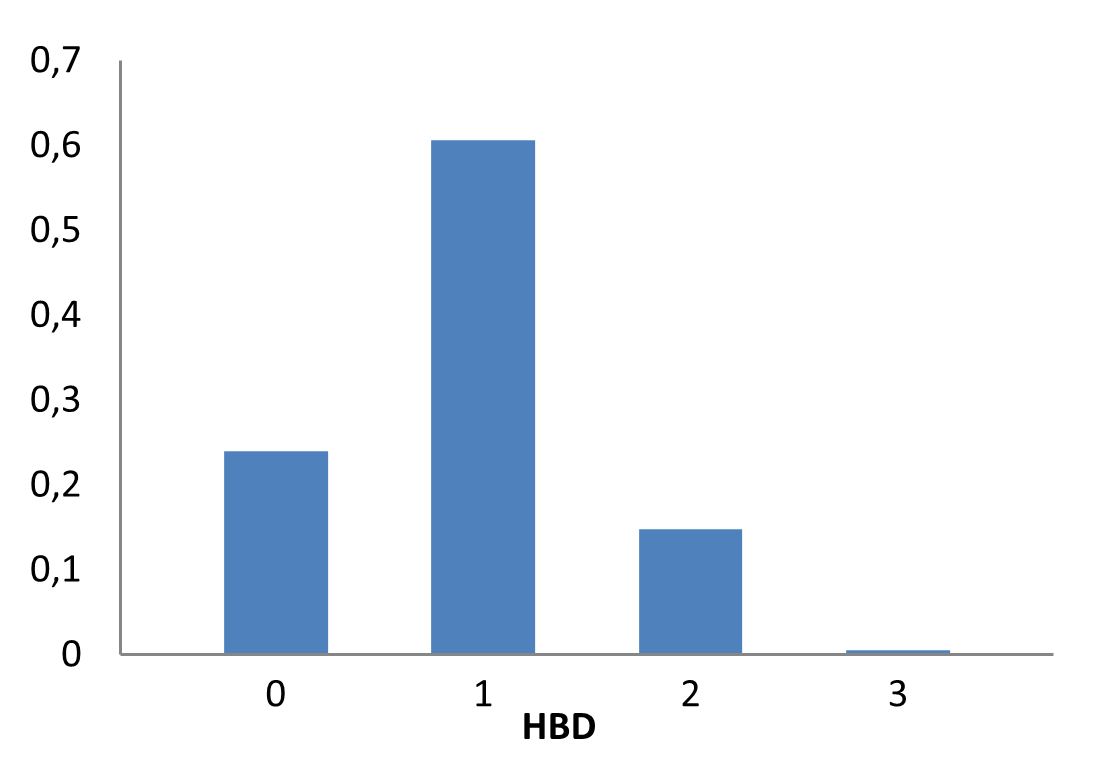Expanding Covalent Warhead Space via SNAr Chemistry
960 compounds
Cysteine-targeting covalent inhibitors are currently dominated by acrylamide and chloroacetamide warheads. In contrast, interest among medicinal chemists in heteroaromatic electrophiles is steadily increasing. Recently, 2‑Sulfonylpyrimidines have gained particular attention due to their ability to undergo cysteine‑selective covalent modification via a nucleophilic aromatic substitution (SNAr) mechanism under mild, physiologically relevant conditions. These features make 2-sulfonylpyrimidines particularly attractive aromatic warheads for the development of next‑generation targeted covalent inhibitors. To support the MedChem community with easy access to these emerging covalent binders, we have developed a 960-compound Covalent 2-Sulfonylpyrimidine Library, available as 10 mM solutions in DMSO.
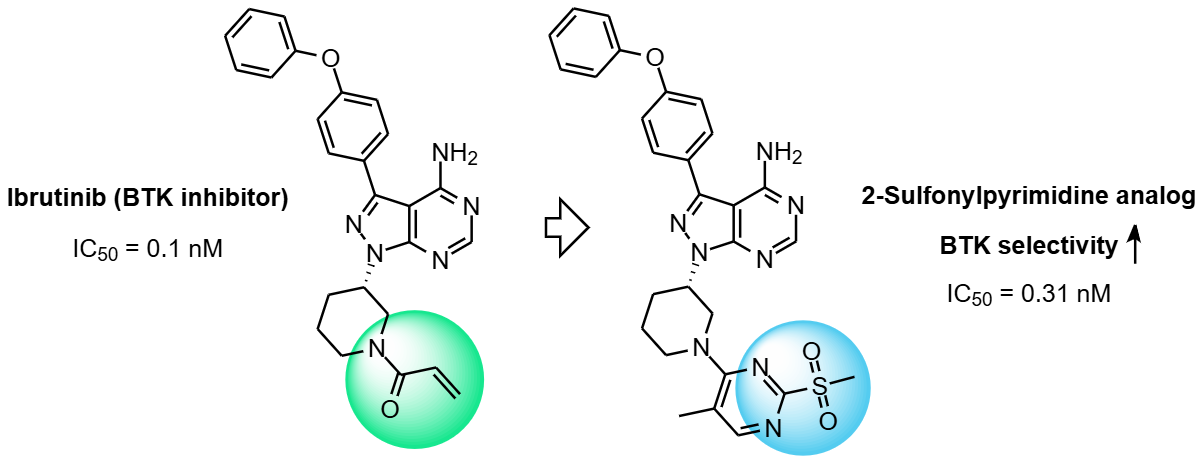
2‑Sulfonylpyrimidine-based Ibrutinib analogs have already been successfully applied in the development of potent and selective BTK covalent inhibitors. These compounds have been shown to function as effective acrylamide surrogates with comparable cellular potency and, importantly, improved kinome selectivity.
Typical Formats
Catalog No.
CSPL-960-0-Z-2
Compounds
960
1 plate
Amount
≤ 300 nL of 2 mM DMSO solutions
Plates and formats
1536 well, Echo Qualified 001-6969 (LP-0400), first four and last four columns empty, 960 compounds per plate
Price
Catalog No.
CSPL-960-10-Y-10
Compounds
960
3 plates
Amount
10 µL of 10 mM DMSO solutions
Plates and formats
384 well, Echo Qualified LDV microplates 001-12782 (LP-0200), first and last two columns empty, 320 compounds per plate
Price
Catalog No.
CSPL-960-50-Y-10
Compounds
960
3 plates
Amount
50 μL of 10 mM DMSO solutions
Plates and formats
384 well, Greiner Bio-One microplates 781280, first and last two columns empty, 320 compounds per plate
Price
*We will be happy to provide our library in any other most convenient for your project format. Please select among the following our standard microplates: Greiner Bio-One 781270, 784201, 781280, 651201 or Echo Qualified 001-12782 (LP-0200), 001-14555 (PP-0200), 001-6969 (LP-0400) or send your preferred labware. Compounds pooling can be provided upon request.
Key features
- Cysteine-selective SNAr reactivity
- Irreversible covalent binding
- Tuned reactivity
- Hydrolytically stable
Library design
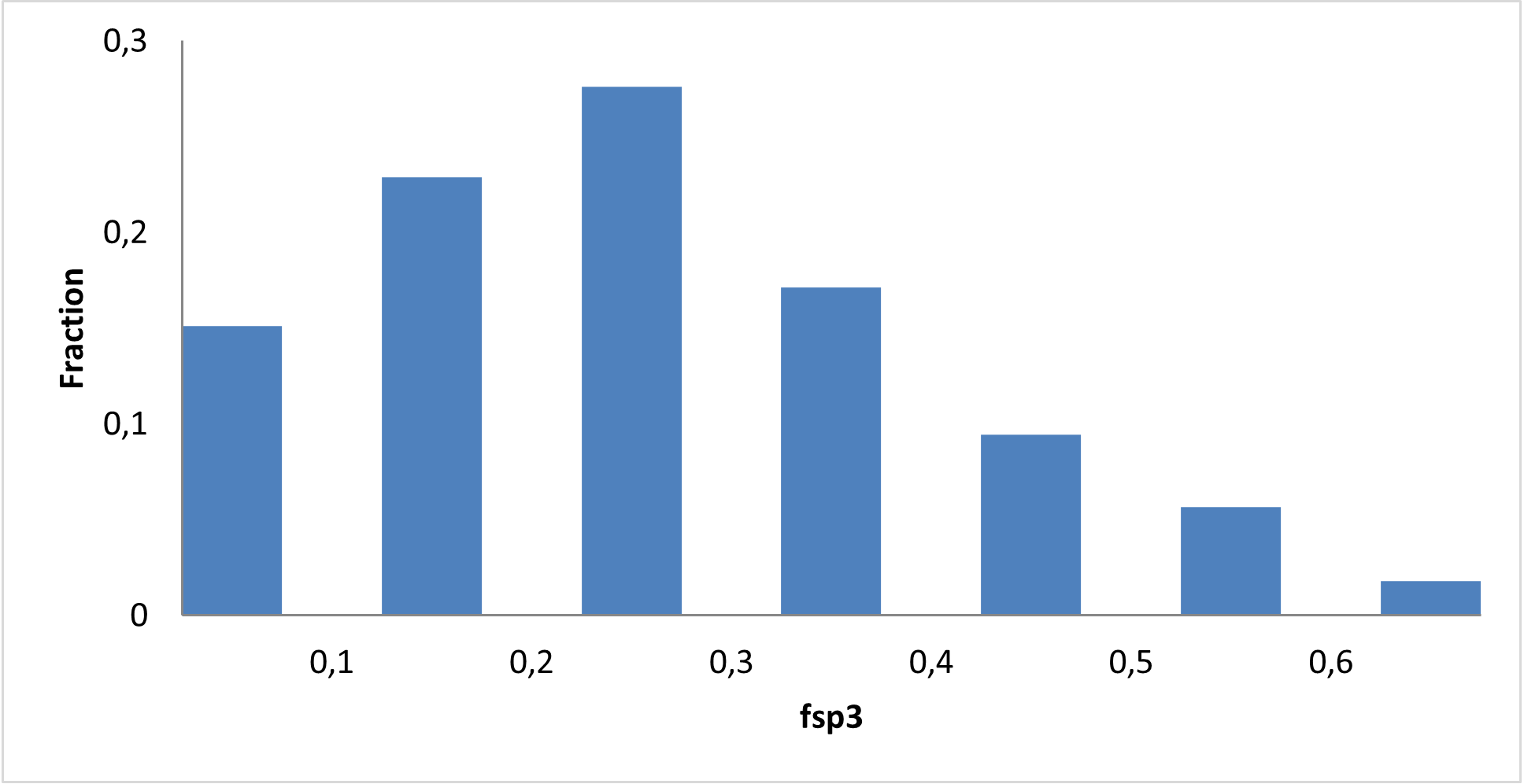
Design of the library focused on functionalization of the 2-sulfonylpyrimidine core using structurally diverse building blocks. Robust synthetic transformations such as arylation and amide bond formation were used to access the final compounds. These synthetic routes provide efficient access to analogs via parallel chemistry. To modulate SNAr electrophilic reactivity, we introduced various electron-donating and electron-withdrawing substituents on the pyrimidine core.
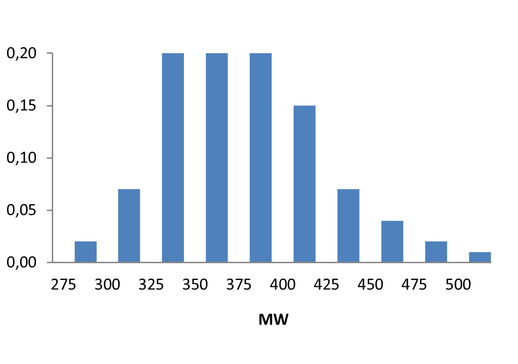
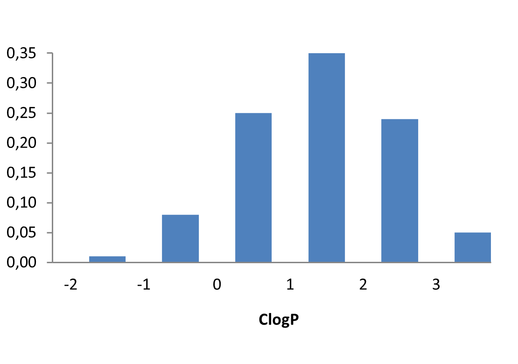
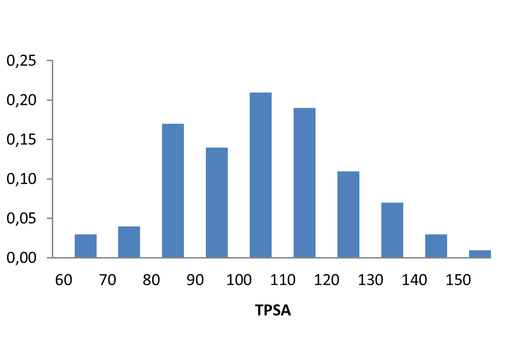
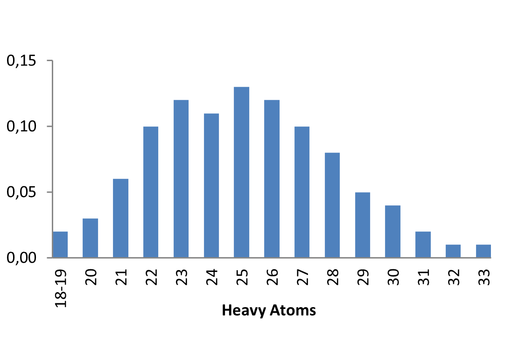
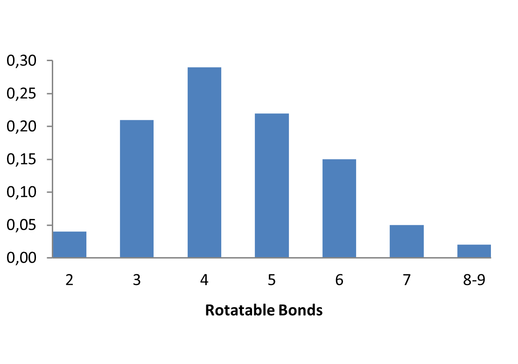
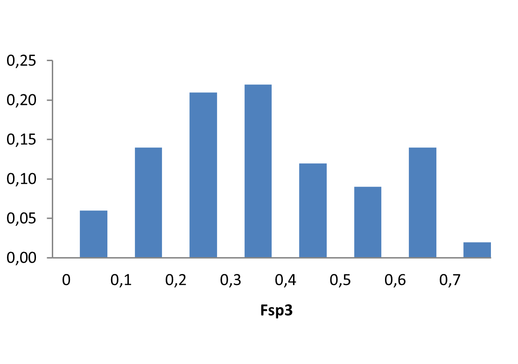
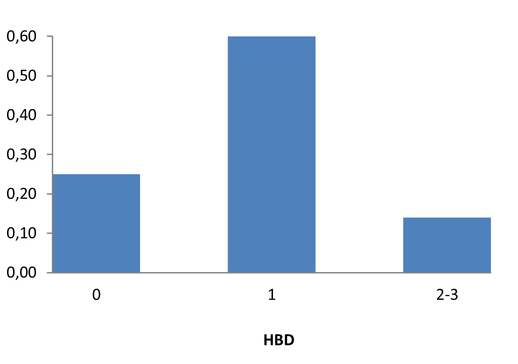
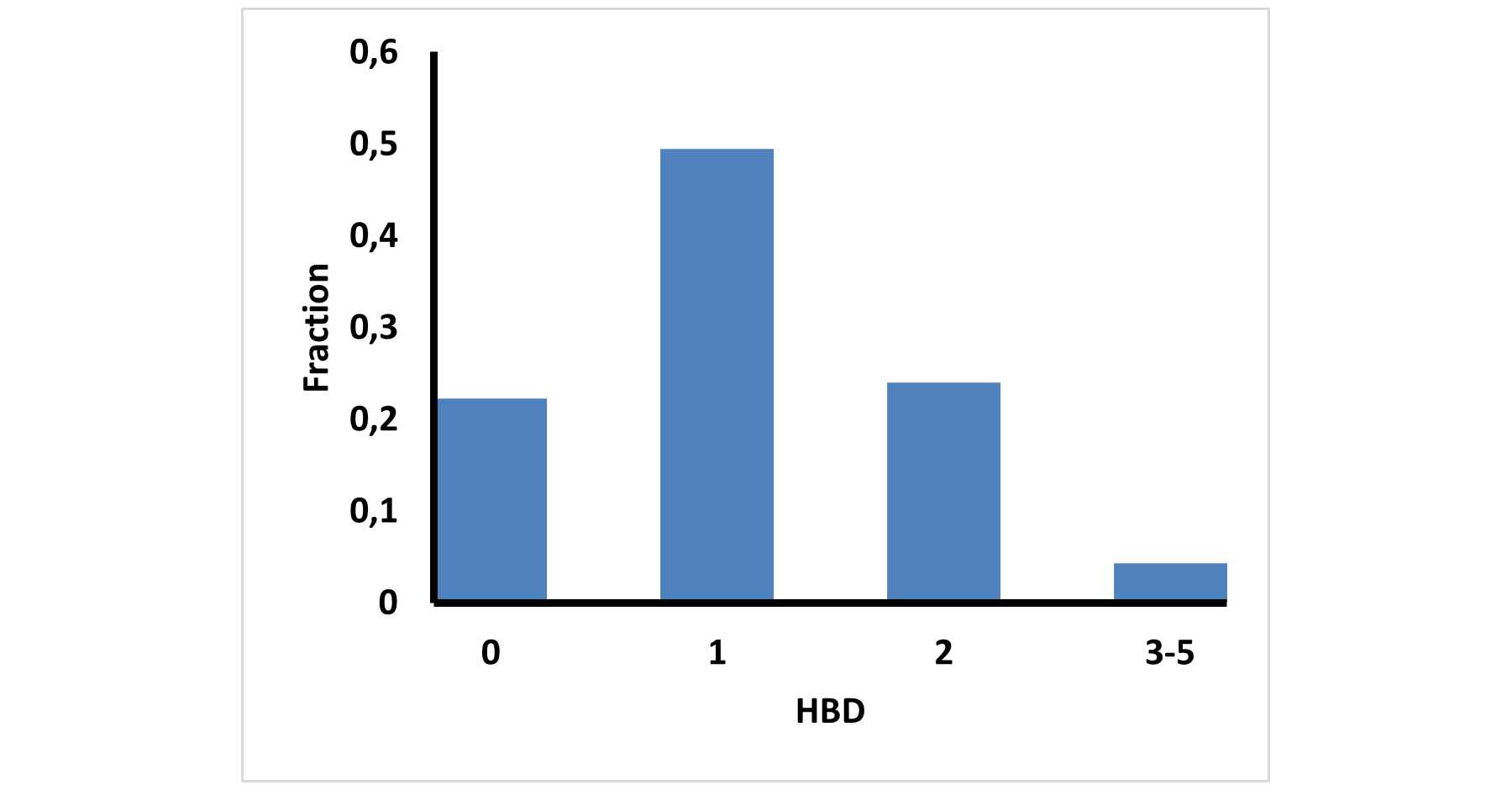
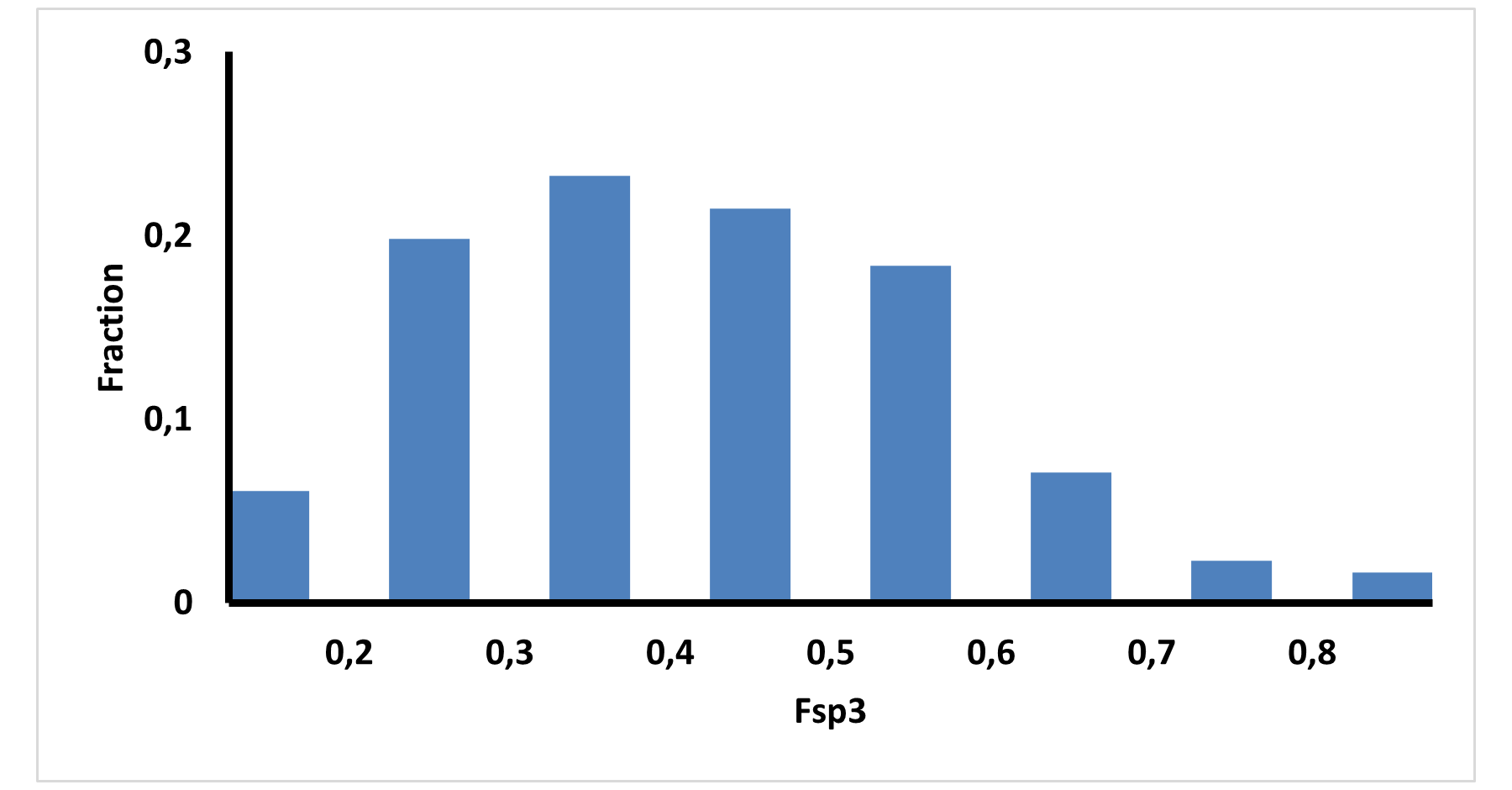
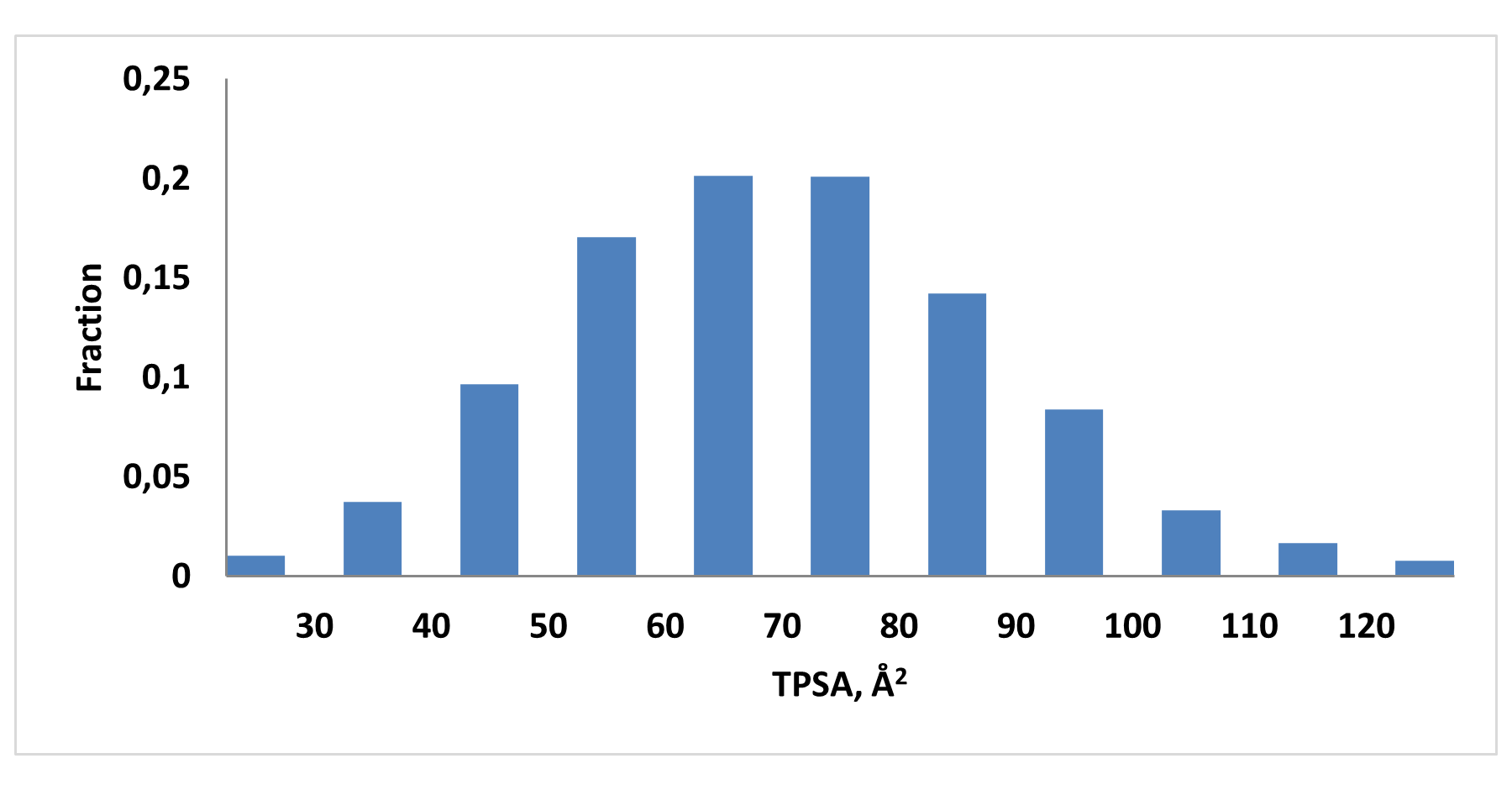
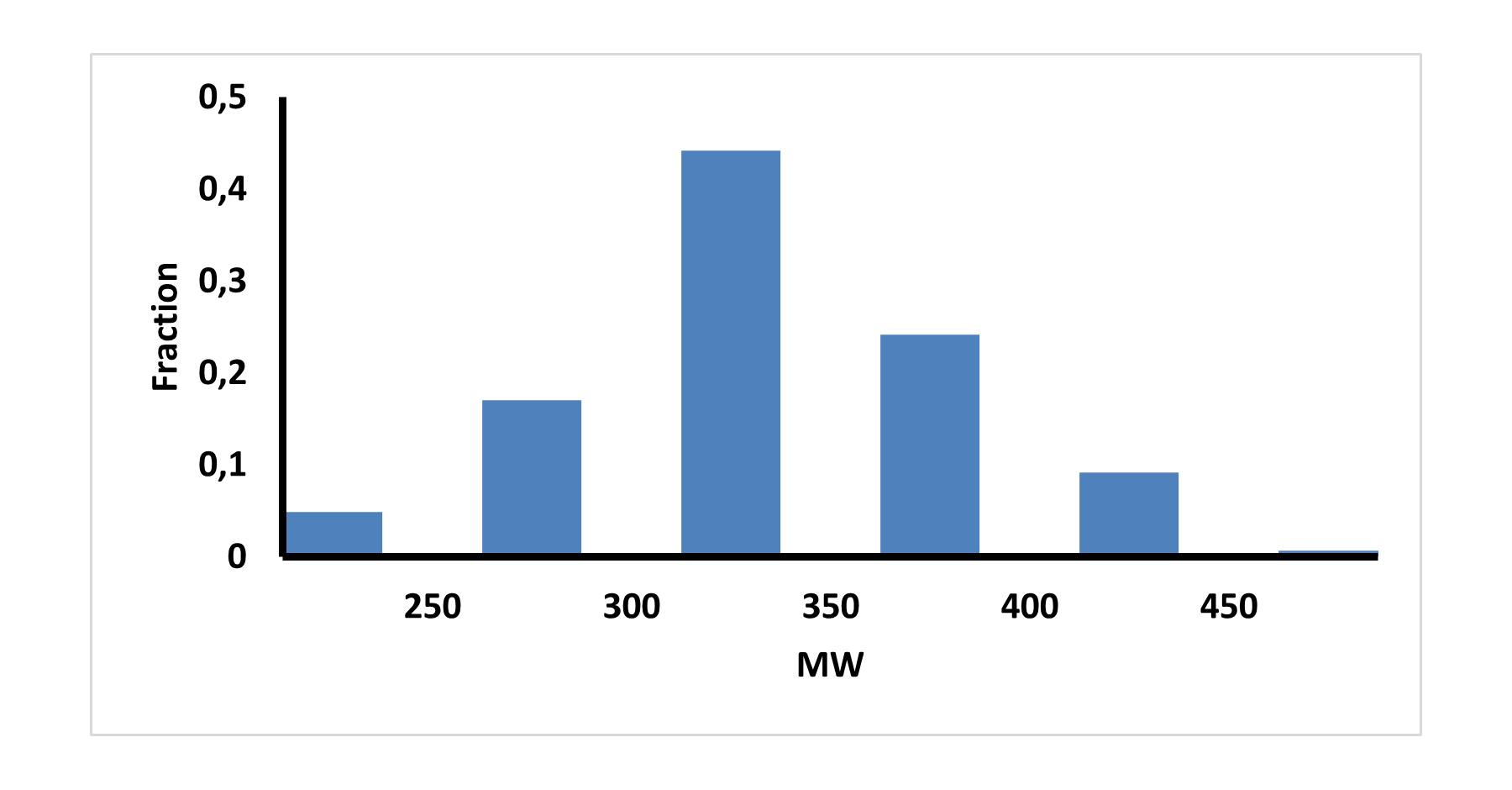
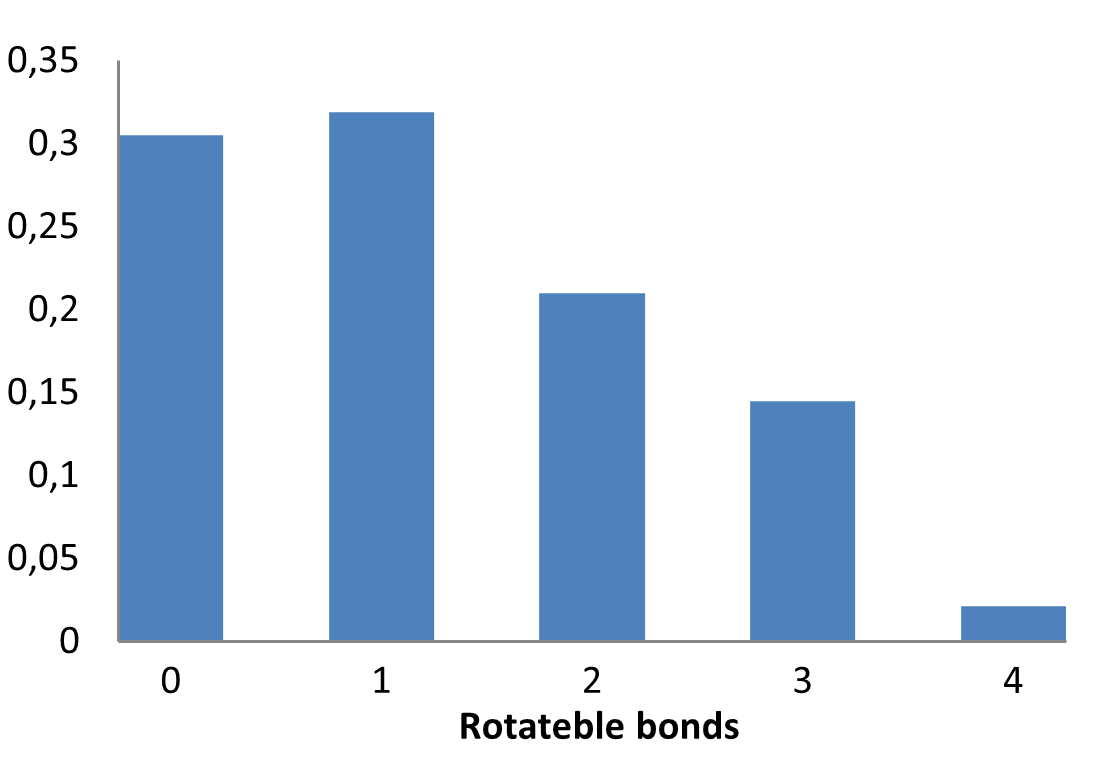
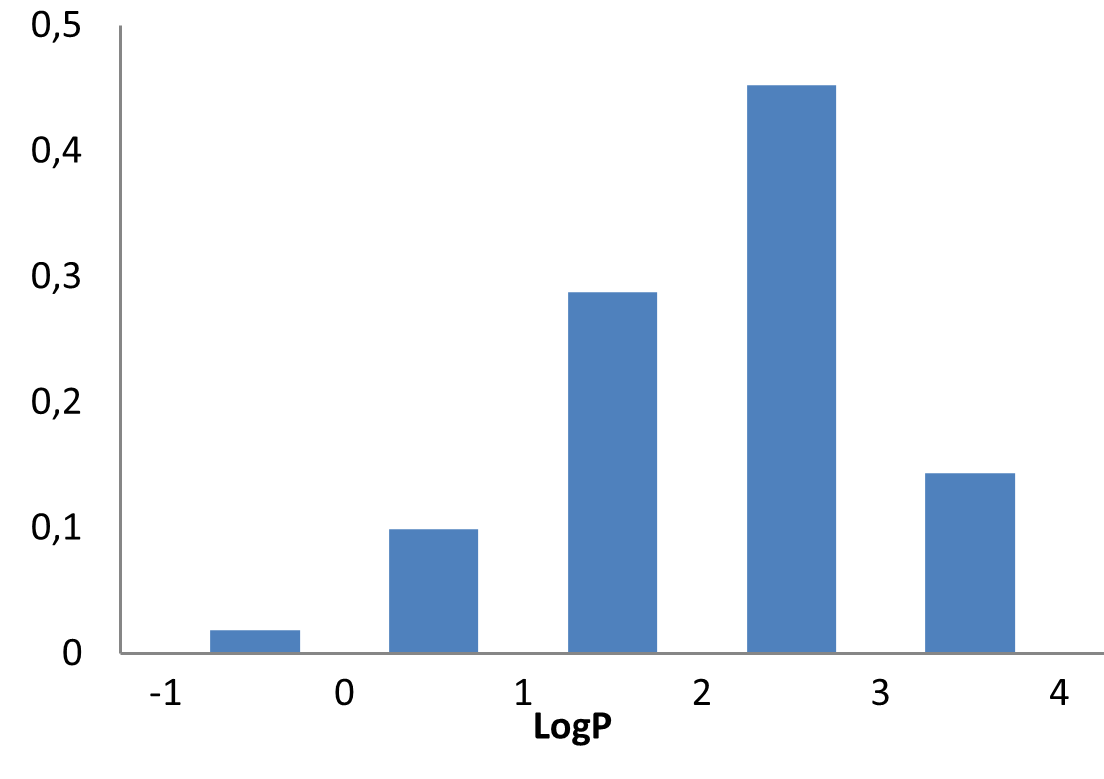
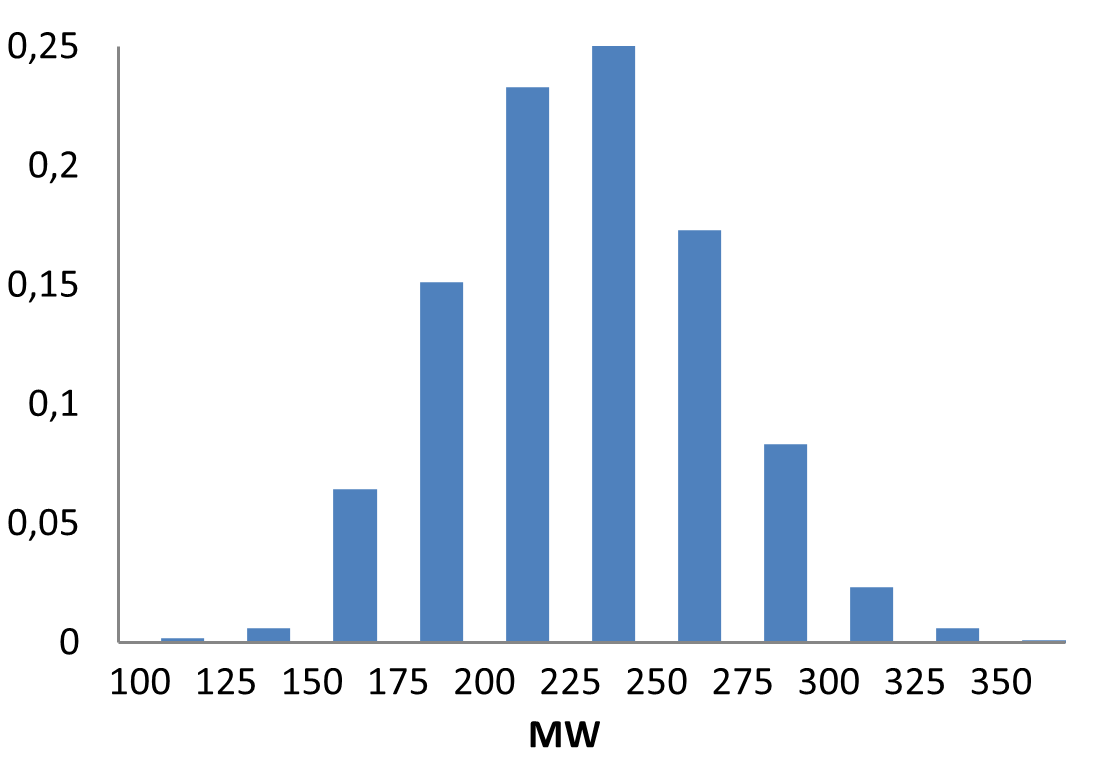
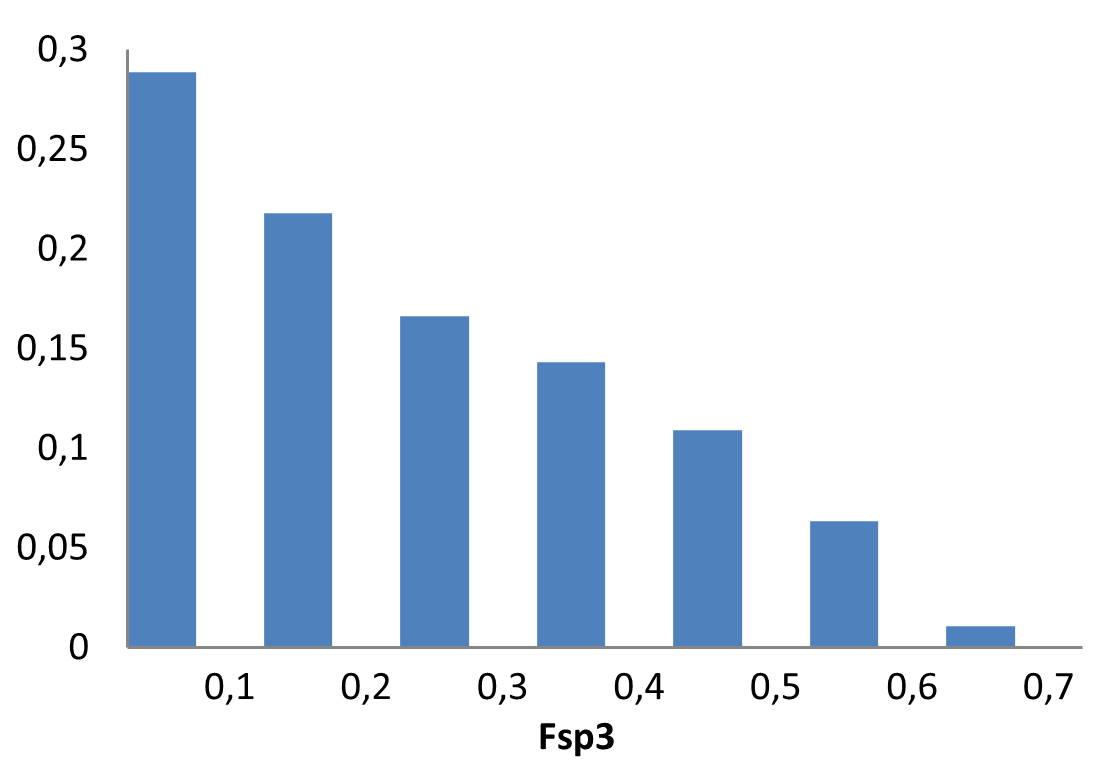
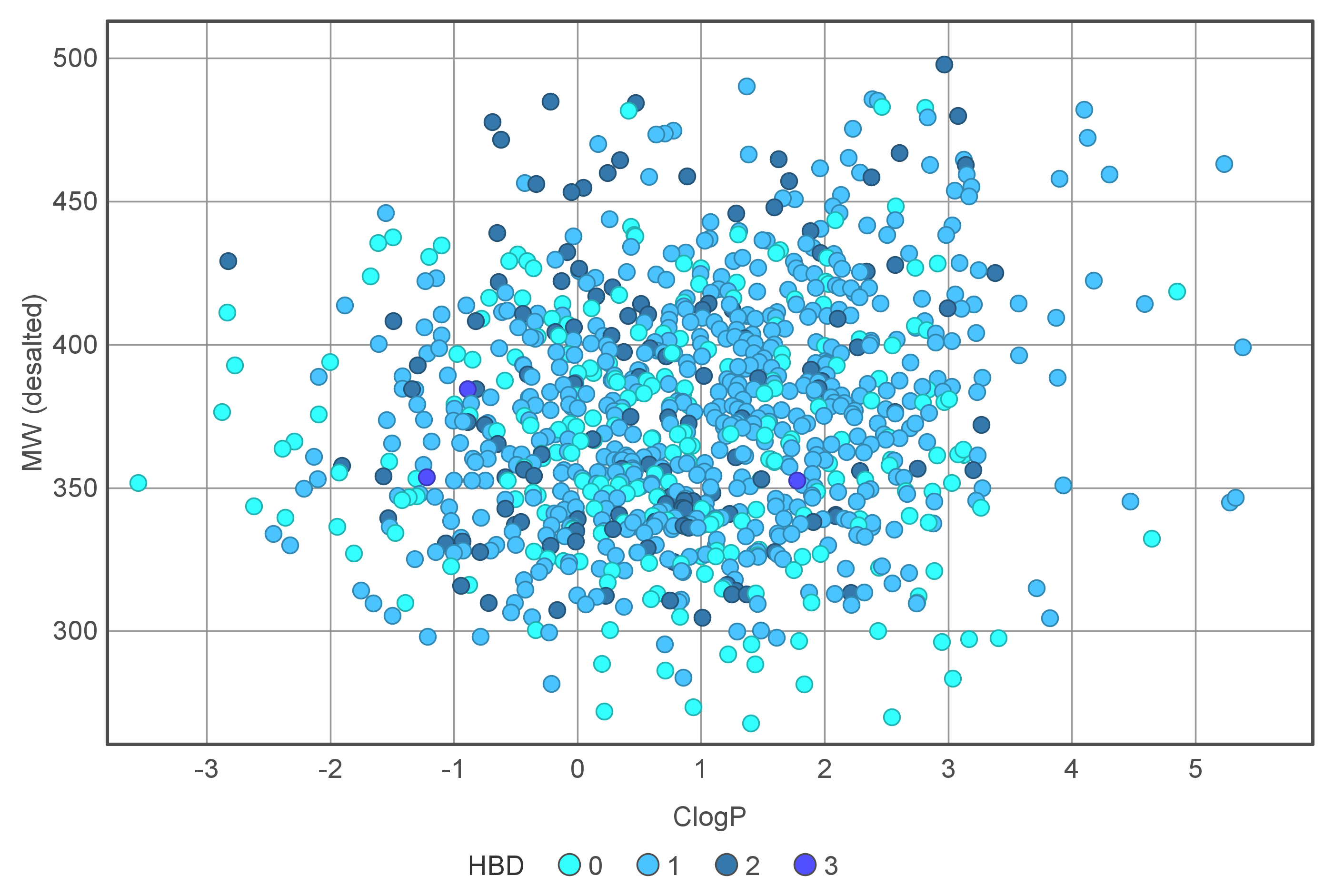
Medicinal chemistry filters were applied during library design to remove undesirable motifs and optimize drug-like properties. Final compound selection for the pre-plated library ensured balanced coverage across key physicochemical parameters (MW, LogP, HBD etc), supporting broad applicability in screening campaigns.
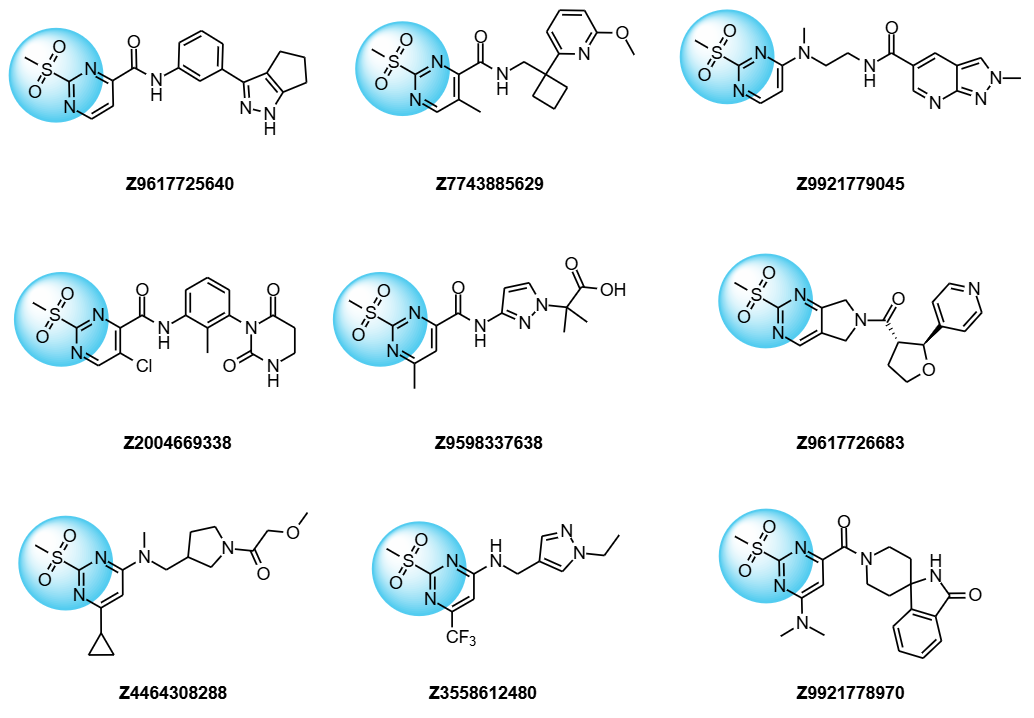
Examples of compounds from Covalent 2-Sulfonylpyrimidine Library
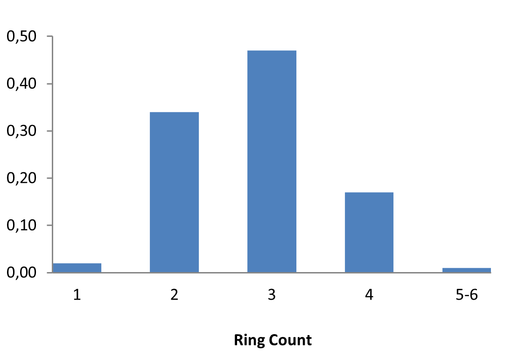
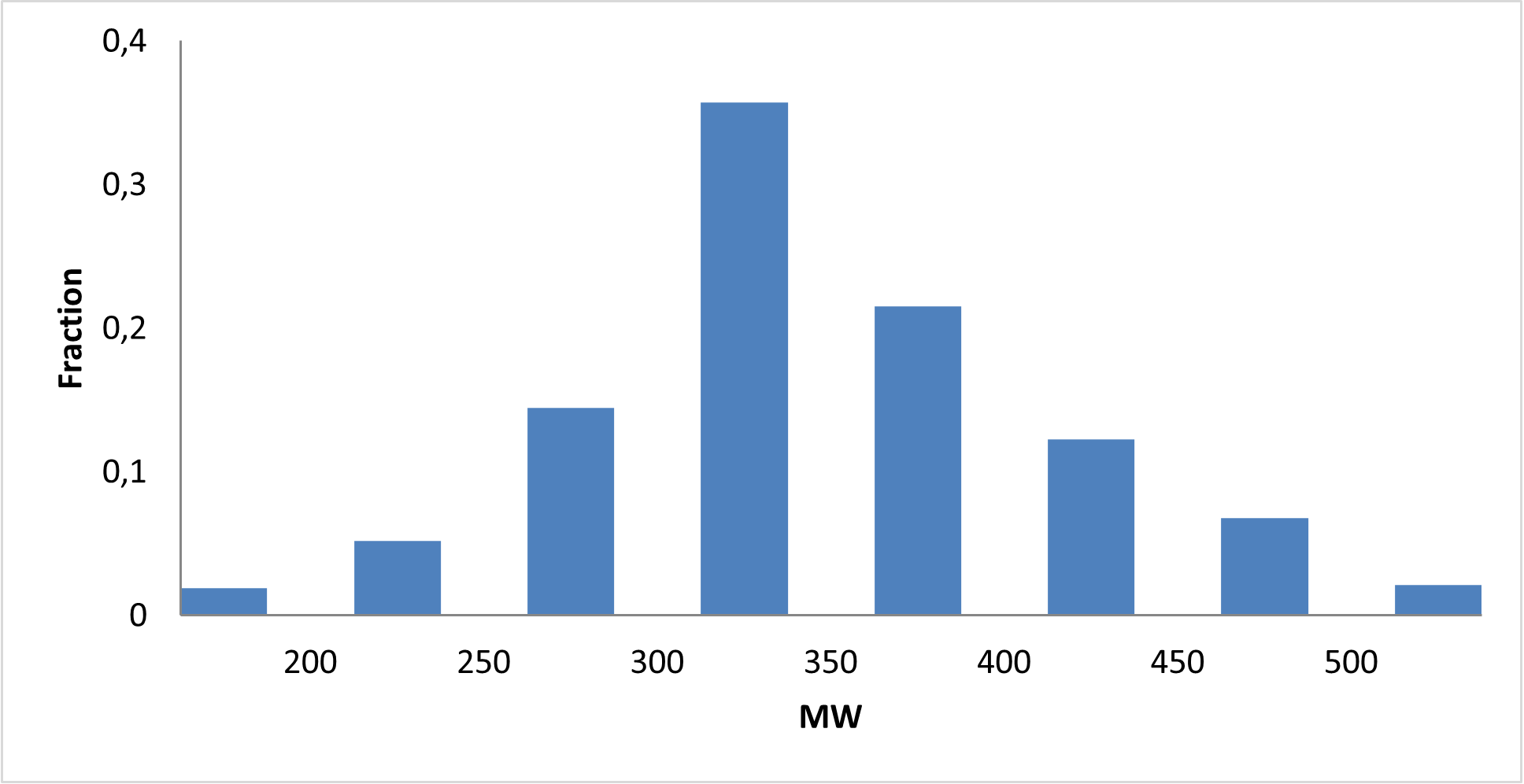
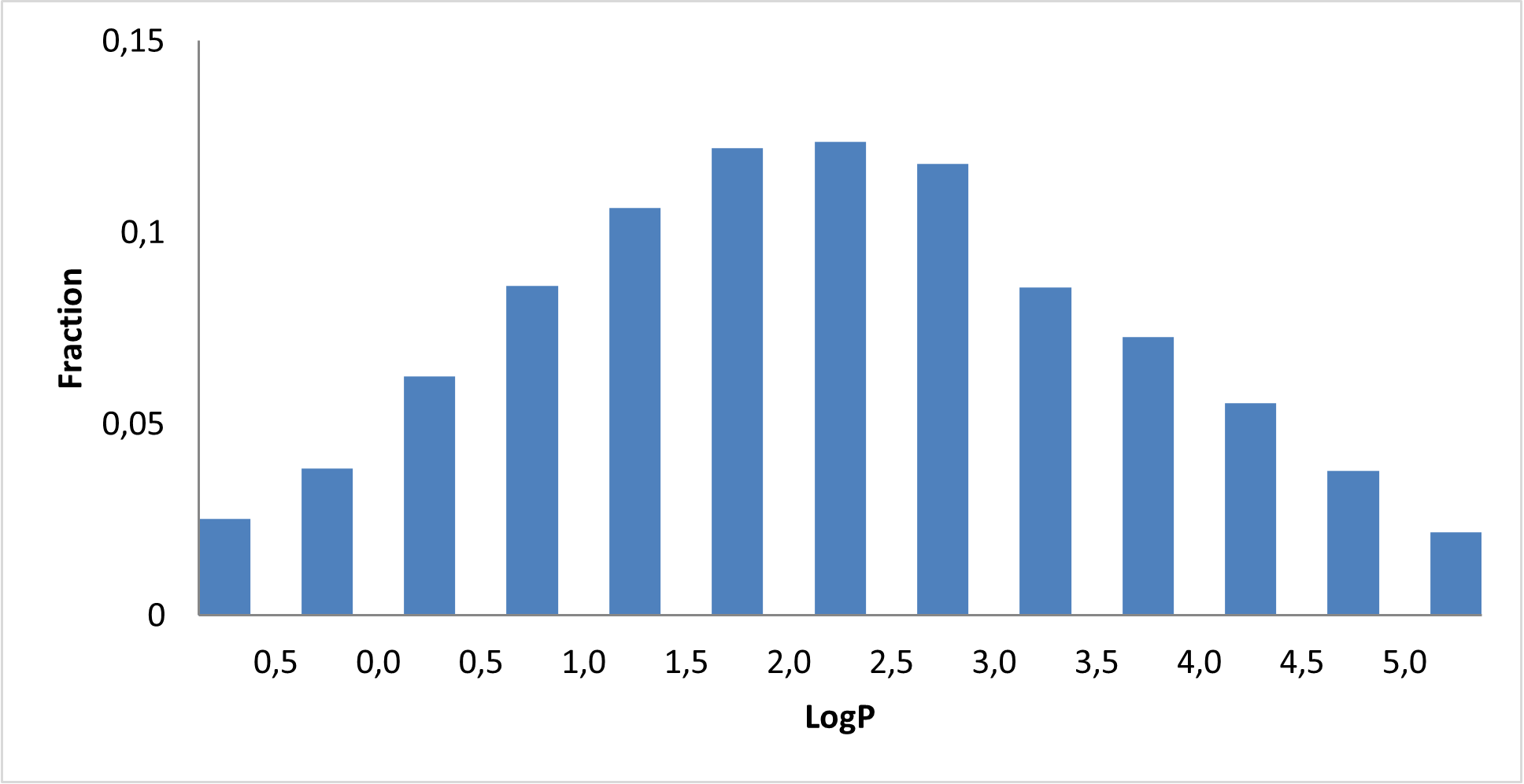
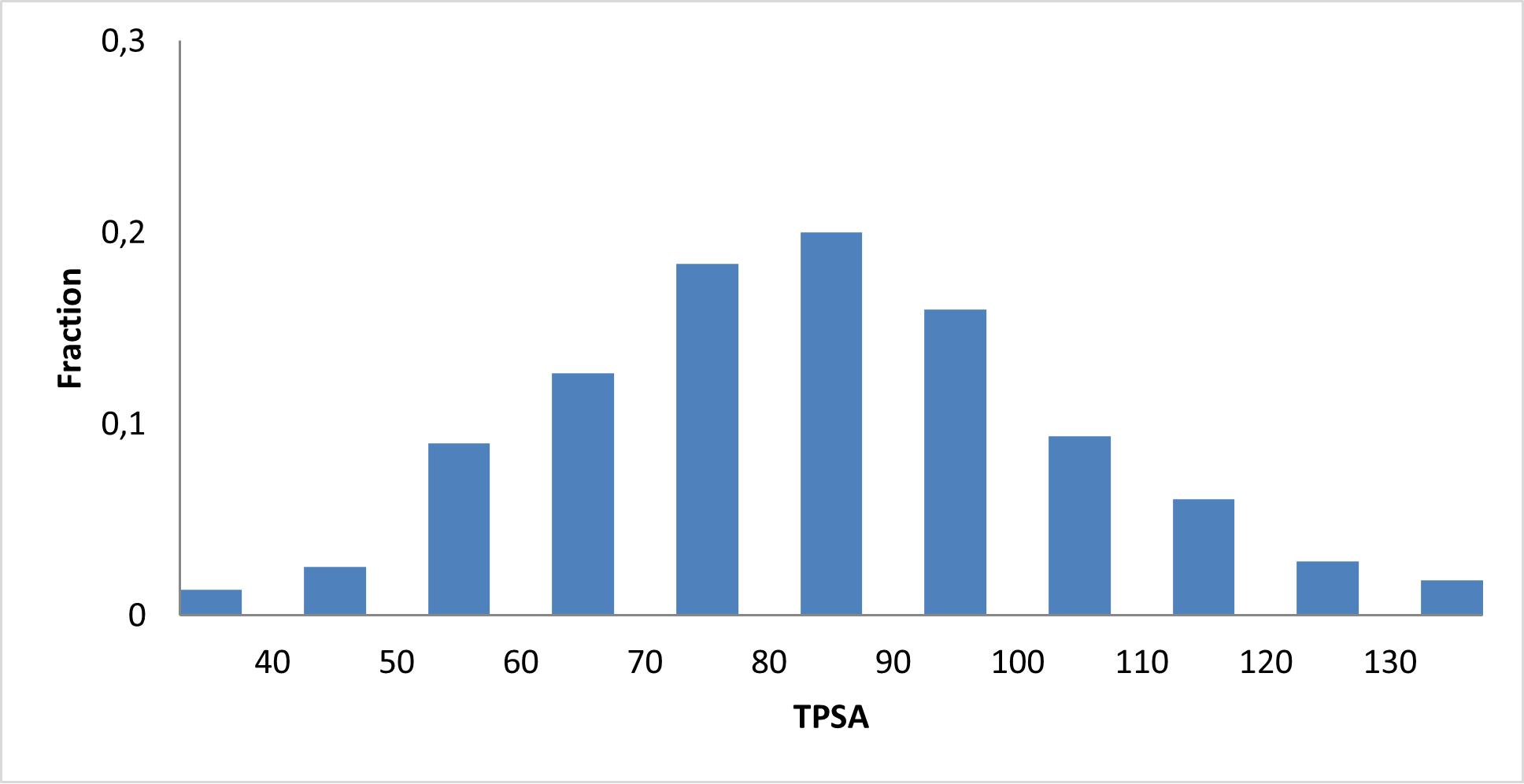
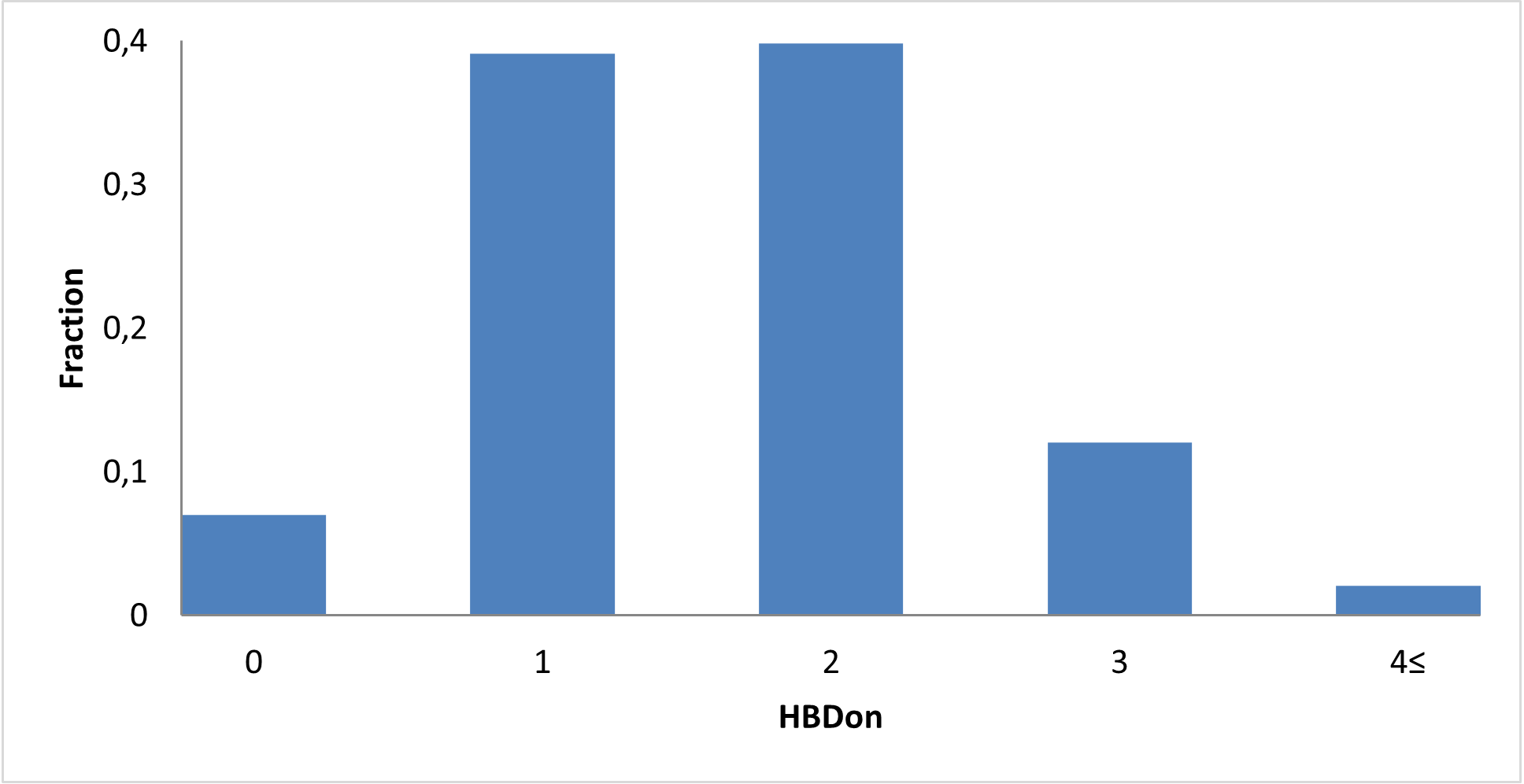
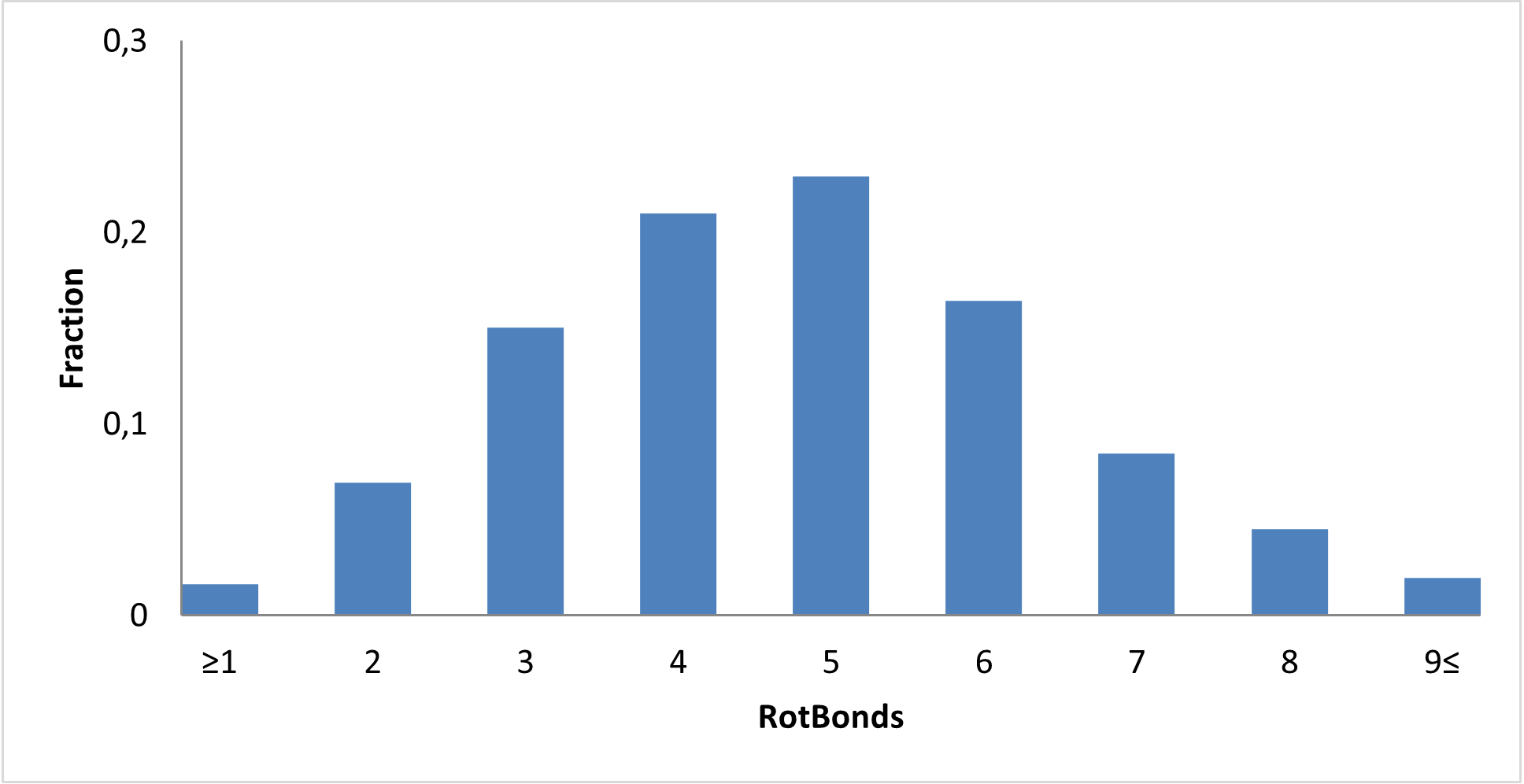
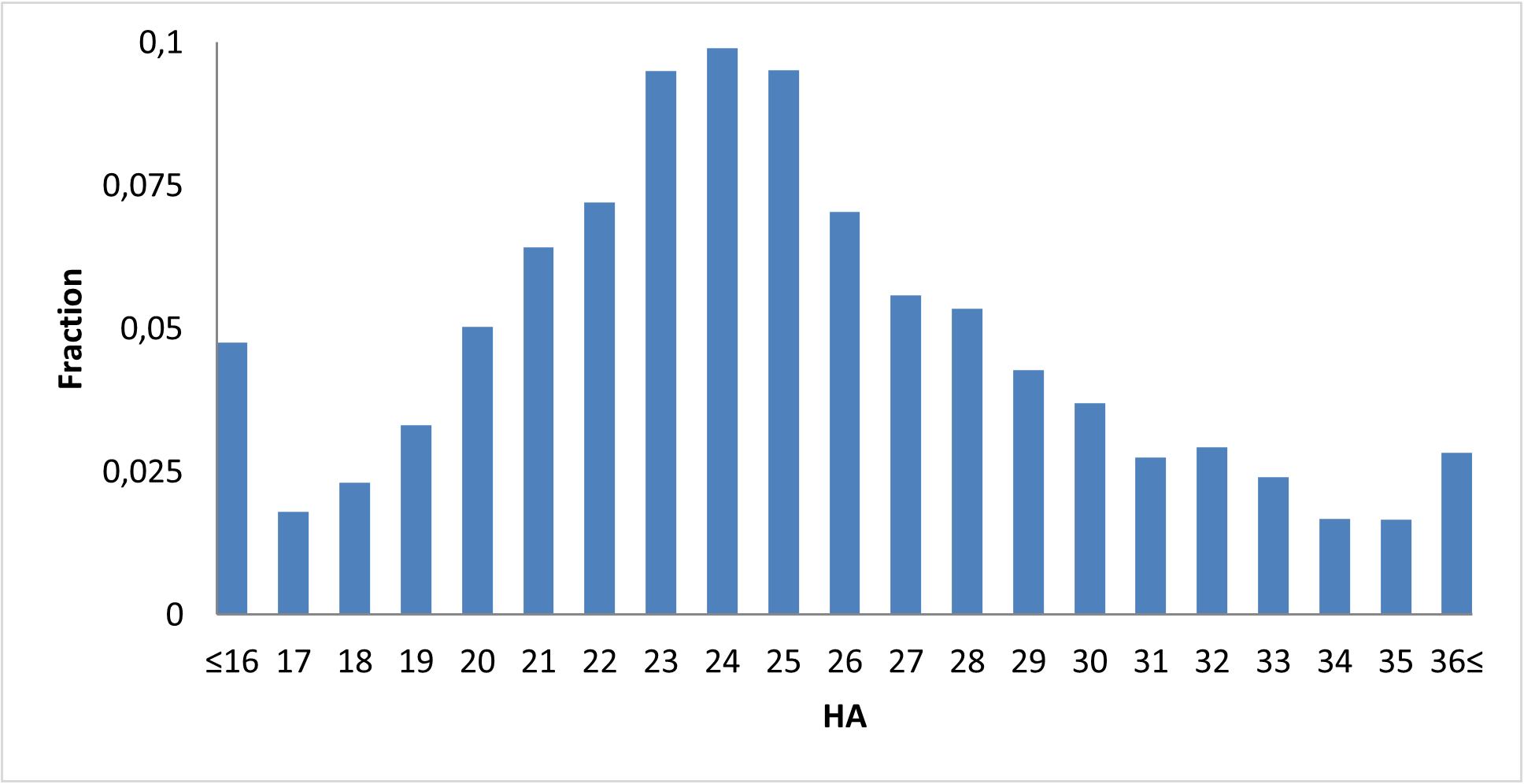
Covalent 2-Sulfonylpyrimidine Library cLogP/MW/HBD distribution
Designed to disrupt c-Myc, built to accelerate breakthroughs in cancer
8 160 compounds
The c‑Myc/Max heterodimer is a basic helix–loop–helix leucine zipper (bHLHZip) transcription factor complex that regulates fundamental cellular processes, including growth, differentiation, metabolism, and programmed cell death. Dysregulation of c‑Myc expression is one of the most characteristic oncogenic events in human biology: c‑Myc is constitutively and aberrantly overexpressed in more than half of all human cancers. Its persistent activity promotes uncontrolled proliferation, maintains stem‑cell‑like self‑renewal, and suppresses senescence and differentiation pathways.
Despite its central role in tumorigenesis and decades of research, no clinically approved therapies currently exist that directly target the c‑Myc/Max complex. This gap keeps c‑Myc among the most compelling but challenging targets in modern drug discovery, with relevance spanning oncology, proliferative diseases, and metabolic disorders.
We assembled a curated c‑Myc‑focused library designed to provide high‑quality, mechanism‑relevant starting points for developing novel c‑Myc modulators and chemical probes.
Typical Formats
Catalog No.
MYC-8160-0-Z-10
Compounds
8 160
7 plates
Amount
≤ 300 nL of 10 mM or lower DMSO solutions
Plates and formats
1536 well, Echo Qualified 001-6969 (LP-0400), first four and last four columns empty, 1280 compounds per plate
Price
Catalog No.
MYC-8160-10-Y-10
Compounds
8 160
26 plates
Amount
10 µL of 10 mM DMSO solutions
Plates and formats
384 well, Echo Qualified LDV microplates 001-12782 (LP-0200), first and last two columns empty, 320 compounds per plate
Price
Catalog No.
MYC-8160-50-Y-10
Compounds
8 160
26 plates
Amount
50 μL of 10 mM DMSO solutions
Plates and formats
384 well, Greiner Bio-One microplates 781280, first and last two columns empty, 320 compounds per plate
Price
*We will be happy to provide our library in any other most convenient for your project format. Please select among the following our standard microplates: Greiner Bio-One 781270, 784201, 781280, 651201 or Echo Qualified 001-12782 (LP-0200), 001-14555 (PP-0200), 001-6969 (LP-0400) or send your preferred labware. Compounds pooling can be provided upon request.
Download SD file
Library design
To build a rationally designed compound library against c‑Myc‑related mechanisms, we focused on three complementary strategies:
- Direct Modulation of the c‑Myc/Max Complex
- Targeting the c‑Myc Promoter G‑Quadruplex
- Indirect Inhibition Through Epigenetic and Co‑Regulatory Targets
Although a liganded structure of the c‑Myc/Max complex is still unavailable, several studies have reported small molecules capable of perturbing its function. Leveraging published models and experimentally validated active compounds, we designed two docking models for direct inhibition:
Model 1: Restriction of c‑Myc/Max flexibility near the E‑box binding pocket. Based on the paper J.Med.Chem., we used structural insights surrounding the compound 4DA (JKY‑2‑169), which binds near the c‑Myc/Max–DNA interface and reduces transcriptional activity by limiting conformational dynamics required for DNA recognition.
Model 2: Direct engagement within the E‑box binding interface. Using the concepts reported in Heliyon and Cell Commun Signal, we incorporated the binding mode of XYA1353, a ligand reported to occupy the DNA‑interaction surface of the c‑Myc/Max heterodimer, thereby preventing formation of the transcriptionally active protein–DNA complex.
These two models represent distinct but complementary approaches for identifying modulators that destabilize or prevent c‑Myc/Max engagement with genomic E‑box sequences.
Given the difficulty of drugging intrinsically disordered transcription factors, promoter‑level regulation has emerged as an alternative strategy. A well‑characterized G‑quadruplex structure within the guanine‑rich region of the MYC promoter can repress transcription when sufficiently stabilized. Therefore, small molecules that bind to or stabilize this G‑quadruplex may indirectly suppress c‑Myc activity.
We incorporated major structural features of known modulators: macrocycle–quadruplex complexes, four- and three-fused aromatic rings, and ligands with two linked aromatic rings.
By mapping these structural features, we sought to capture molecules capable of stabilizing the G‑quadruplex and thereby suppressing c‑Myc transcription.
Because c‑Myc/Max relies heavily on chromatin‑associated partners and epigenetic modifiers, indirect inhibition is another promising therapeutic strategy. We focused on targeting key proteins implicated in c‑Myc‑dependent transcriptional regulation: EZH2, HDAC2, KDM4A, KDM4B, KDM4D. These targets are functionally linked to chromatin environments that facilitate c‑Myc–driven gene expression. Modulation of their activity has been shown to alter c‑Myc transcriptional output, making them relevant secondary targets for targeting c‑Myc.
Through these three mechanistic approaches — direct inhibition, promoter G‑quadruplex stabilization, and modulation of c‑Myc‑associated epigenetic regulators — we constructed a diverse and biologically relevant compound library, designed to explore multiple intervention points of c‑Myc regulatory. This multifaceted approach increases the likelihood of identifying promising modulators of the most important oncogenic drivers in human disease.
All protein-ligand complexes were evaluated for stability using a 100 ns MD simulation. The optimized and stable complexes were then used for molecular docking calculations. The following PDB structures have been used for optimization and further simulations – direct binding to c-Myc/Max: 1nkp and 5i50, c-Myc/Max G-quadruplex: 7n7e, 7png, 6jj0, 5w77, and 2a5r, indirect inhibition: 5ls6, 8bpc, 7mos, 7ltk, 7kbh, 6xeb, 6g3o, 8a0b, 7zzw, 7zzt, 5ix0, 3max, 6h4w, 6h4u, 6hgt, 5fpv, 5f3i, 5f3c, 3pdq, 6cg2, 6cg1, 5vmp, 5a7w, 5a7s, 5f5i, 5f39, 3rvh, 6h8p, 7jm5, 5f5c, 5fp4, 5fp8, 6etg, 6ete, 6f5s, 6h0y, and 6h11.
MD simulation result
Model 1
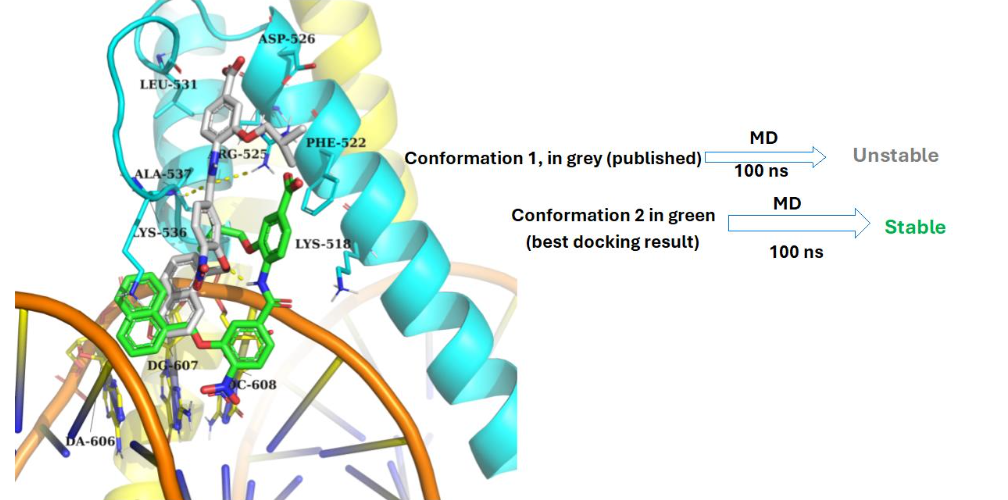
Binding models between c-Myc/Max and 4DA (“JKY-2-169”). DNA in orange, Myc in yellow, Max in cyan.
Model 2
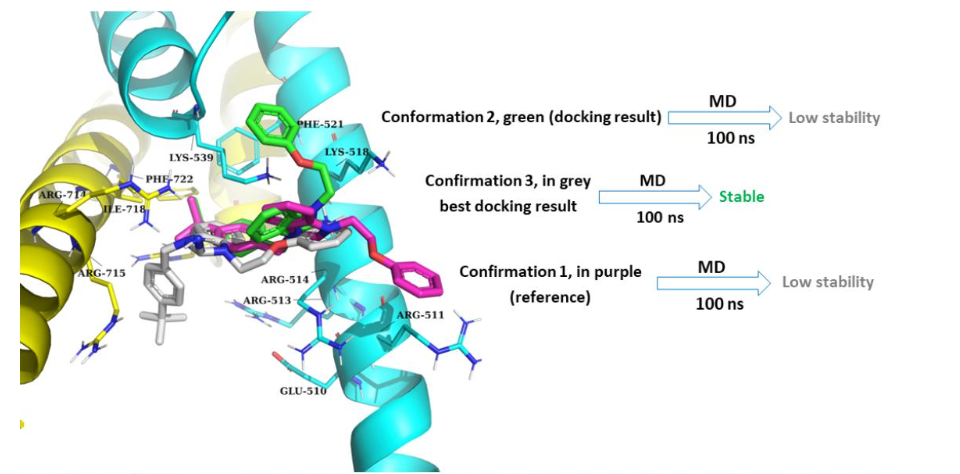
Binding models between c-Myc/Max and XYA1353. DNA in orange, Myc in yellow, Max in cyan.
Examples of molecular docking simulation, and used pharmacophore models, c-Myc/Max protein, direct interaction
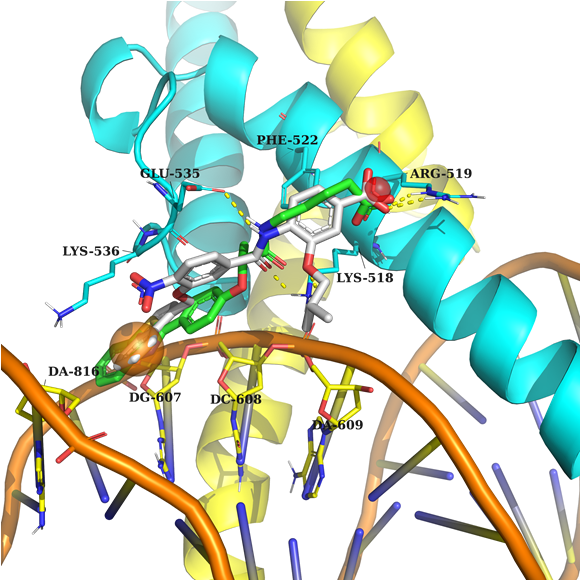
Example 1 (PDB ID 1nkp, c-Myc/Max)
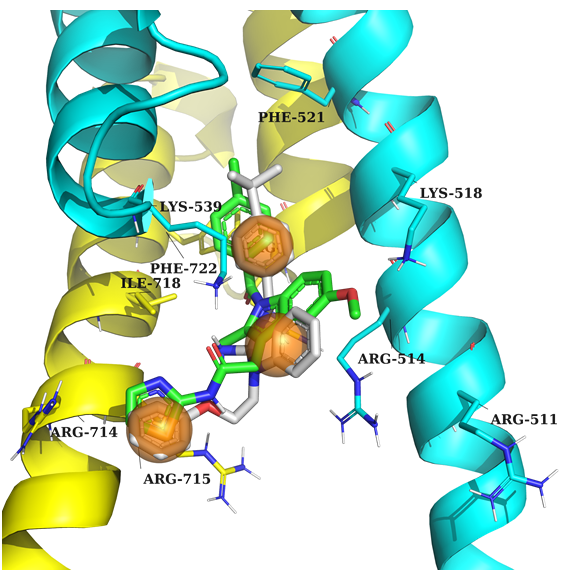
Example 2 (PDB ID 1nkp, c-Myc/Max)
Example 1: According to the pharmacophore model, the ligand's acceptor interacts with Arg 519, and the aromatic ring fills the sub-pocket created by Lys 536 and the nucleotide environment of DNA.
Example 2: Key feature – ligand should contain 3 aromatic rings to create stacking interaction with nearby amino acids.
DNA is in orange, Myc in yellow, Max in cyan, native ligand in grey, docked example in green.
Examples of molecular docking simulation, and used pharmacophore models, c-Myc/Max G-quadruplexes
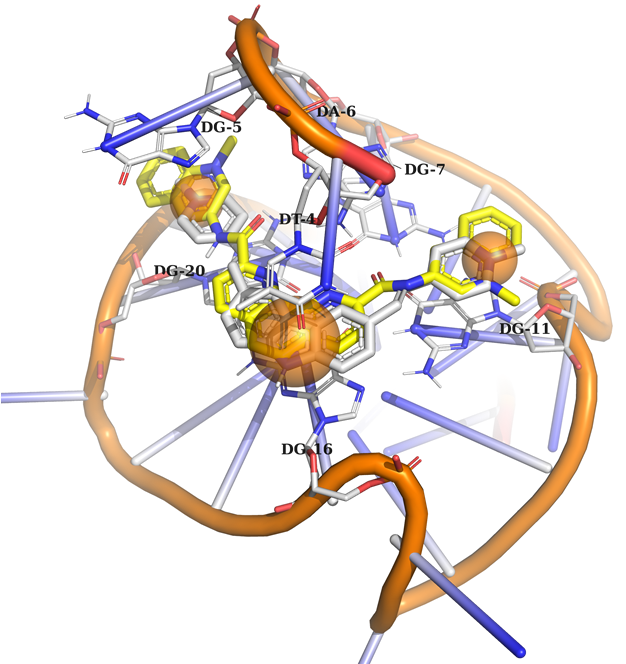
Example 1 (PDB ID 6jj0, c-MYC G-quadruplex)
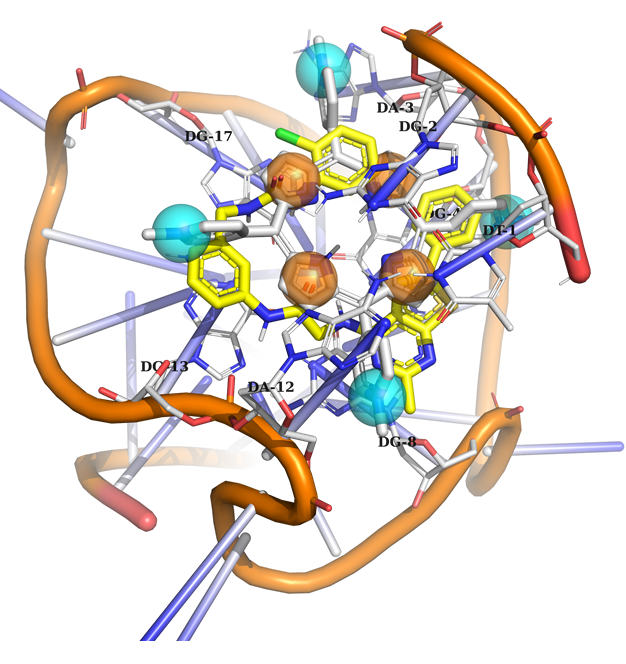
Example 2 (PDB ID 2a5r, c-MYC G-quadruplex)
Example 1: Ligand should contain 3 aromatic rings to create a stacking interaction with the nucleotides.
Example 2: The model contains 8 binding points, including 4 aromatic and mixed aromatic or N+ feature. According to this model, a potential ligand must occupy at least 3 out of 8 binding points to be selected.
DNA in orange, native ligand in grey, docked ligand in yellow.
Examples of molecular docking simulation, and used pharmacophore models, Indirect inhibition strategies
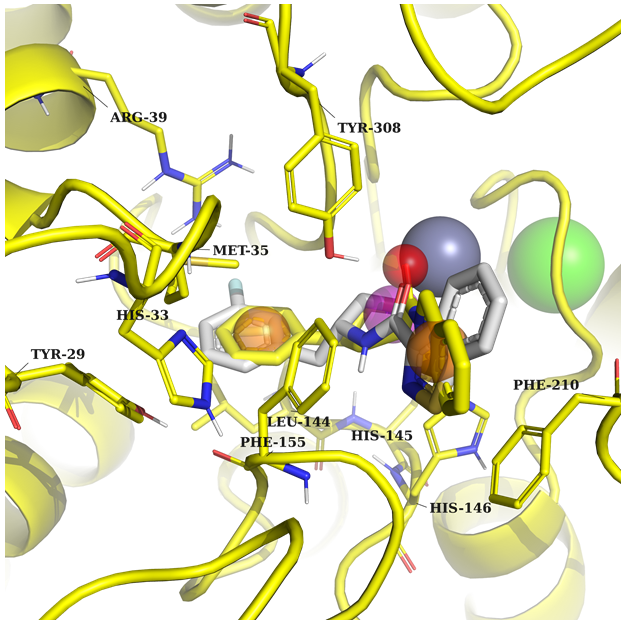
Example 1 (PDB ID 8a0b, HDAC2)
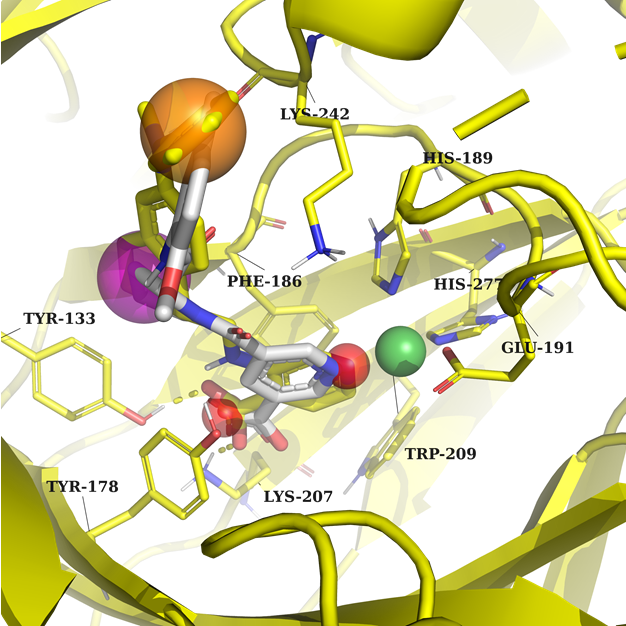
Example 2 (PDB ID 7jm5, KDM4B)
Example 1: A potential ligand requires two aromatics to create π-stacking interaction with Phe155, Phe210, and Tyr308 in one part, and Arg39 and Met35 on the other part of the pocket. Key features: acceptor, ring and donor near the metal atom.
Example 2: Two acceptor features interacting with Lys207, Tyr133, and a metal. Any ring or aromatic ring to fill the binding pockets and create interaction with Lys242.
In both cases, the protein and native ligand are in yellow, and the docked structure in grey.
Innovatively designed and experimentally confirmed library for reliable enantioselective ASMS screening
12 673 compounds
Rapid progress in drug discovery depends on the continuous development of innovative technologies and advanced screening approaches. Tackling challenging targets and so‑called “undruggable” proteins requires not only novel techniques and screening methods but also access to high‑quality, well‑characterized compound libraries.
The recently introduced enantioselective protein affinity selection mass spectrometry (E‑ASMS, Nature Comm. 2025) approach streamlines the time needed for hit identification and hit validation. This method accelerates early drug discovery by enabling the detection of weak binders, delivering critical insights into compound selectivity, and providing orthogonal confirmation of binding events — all without reliance on traditional counter‑screen assays. When combined with carefully designed compound library, E‑ASMS offers a powerful platform for fast identification of promising therapeutic leads.
In collaboration with leading experts from top Pharma companies and the Structural Genomics Consortium (SGC), we carefully designed, experimentally characterized, and validated — through numerous screening campaigns — the first edition of our Enantio‑ASMS Library (E‑ASMS).
Typical Formats
E-ASMS Library is available for supply in various pre-plated, including but not limited to the following most popular ones:
Catalog No.
E-ASMS-12673-Y-10
Compounds
12 673
40 plates
Amount
10 µL of 10 mM DMSO solutions
Plates and formats
384-well echo-source plates, 320 compounds per plate, first two and last two columns empty
Price
Catalog No.
E-ASMS-12673-Y-25
Compounds
12 673
40 plates
Amount
25 µL of 10 mM DMSO solutions
Plates and formats
384-well plates, 320 compounds per plate, first two and last two columns empty
Price
Catalog No.
E-ASMS-12673-X-50
Compounds
12 673
159 plates
Amount
50 µL of 10 mM DMSO solutions
Plates and formats
96-well plates, 80 compounds per plate, 1 and 12 columns empty
Price
*Please select among the following standard microplates: Greiner Bio-One 781270, 784201, 781280, 651201 or Echo Qualified 001-12782 (LP-0200), 001-14555 (PP-0200), 001-6969 (LP-0400), C52621 or send your preferred labware. Compounds pooling can be provided upon request.
Plated library in stock
Library Design
E-ASMS Library has been developed to accelerate the early stages of drug discovery by providing a diverse and well-characterized set of small molecules. Each compound has been selected to have only one chiral atom and be a mixture of only two enantiomers, which can be easily separated by using one of four standard and described in the paper conditions.
Additionally, the library is designed to maximize chemical space coverage, maintain drug‑like properties, and support efficient hit identification across diverse therapeutic areas. Rigorous quality control and SFC chiral separation of each compound ensures reproducibility and reliability, making this library a robust resource for screening campaigns.
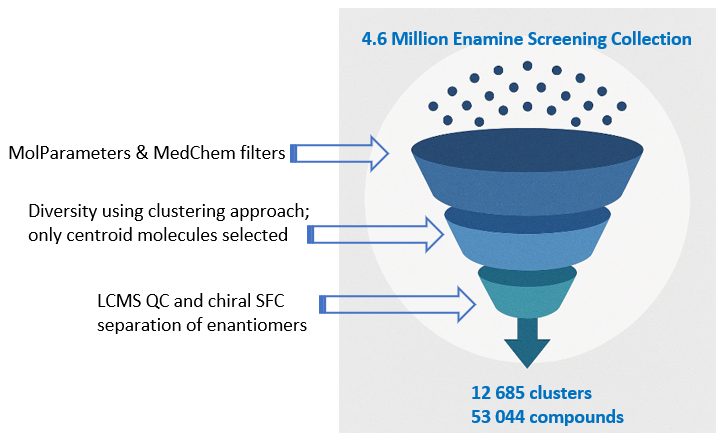
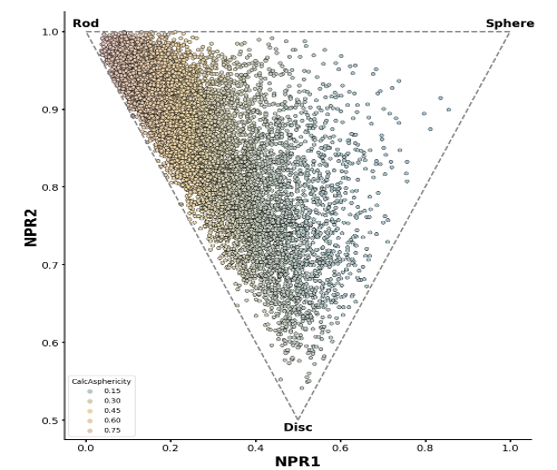
UMap projection to the entire enamine stock collection of 4.6 million compounds and PMI plot.

Each compound in the library is supported with pre-calculated analogs from stock and from REAL Space. Links to the files with calculated analogs are available from SD file in a separate column.
Analogs calculation and search in REAL Space
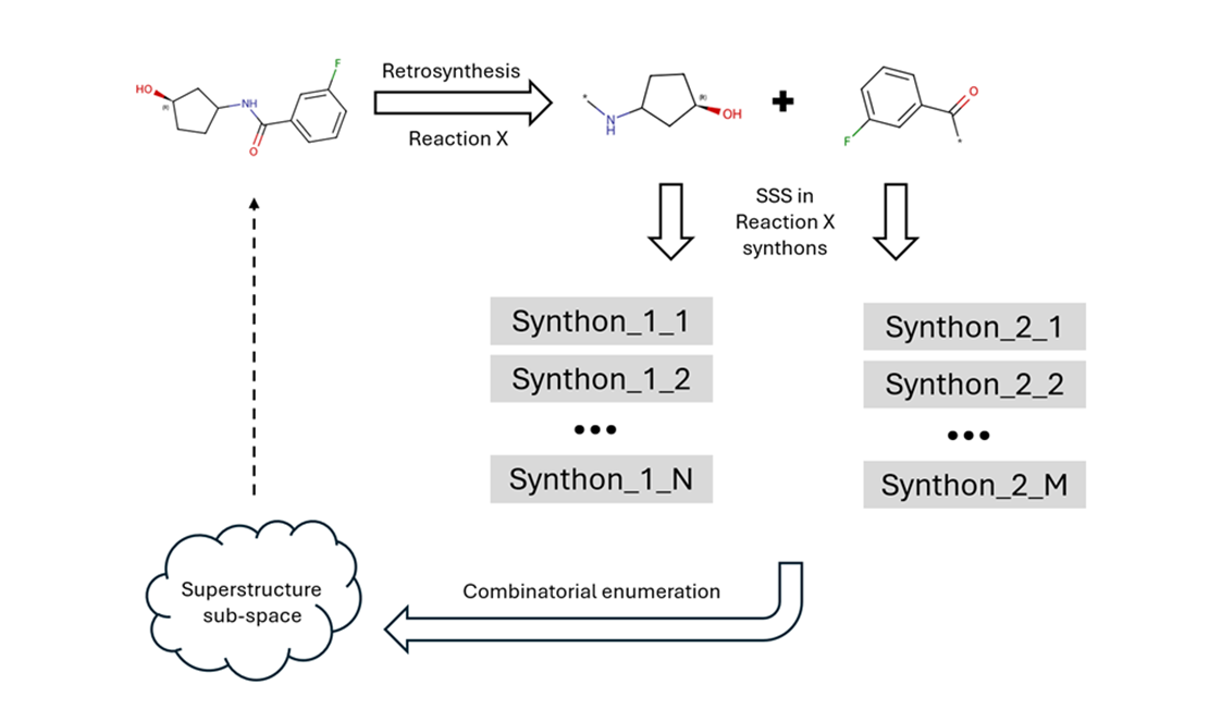
1. Reaction guided search of analogs
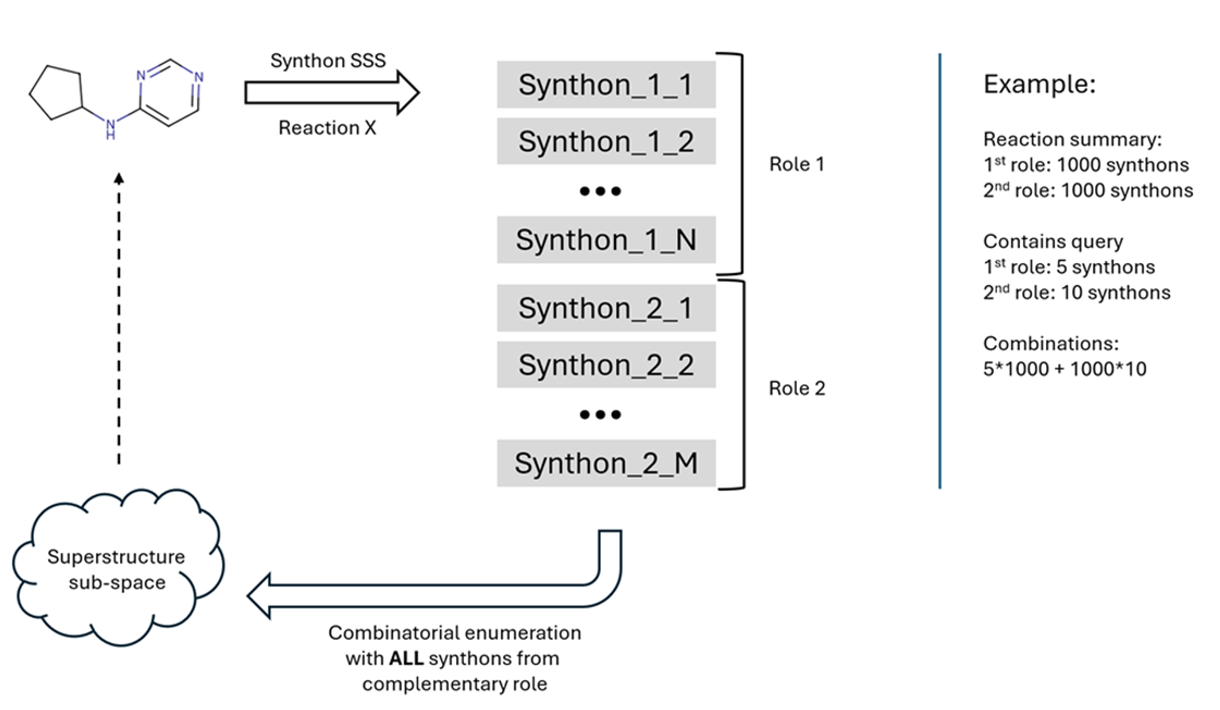
2. Synthon-based sub-structure search
We invite medicinal chemists, biologists, and drug hunters to integrate the E-ASMS Library into their discovery pipelines. By leveraging this curated library, researchers can rapidly identify novel scaffolds, explore structure activity relationships, and generate data that informs lead optimization process. Contact us today to access the library and accelerate your next breakthrough.
Support
We offer comprehensive support in developing your hit compounds. Naturally such programs are realised most efficiently when biological actives originate from our screening collection. However, even if the hit compounds are from the collections of other vendors lead identification and optimization projects can proceed most productively in our hands. Sometimes for this we only need to synthesize first examples of the given chemical series and validate synthesis route.
Most diverse halogen-enriched fragments
1 920 compounds
The concept of Halogen-Enriched Fragment Libraries (HEF) has gained significant traction in medicinal chemistry, supported by multiple successful screening campaigns.[1,2] These libraries are designed to explore halogen-mediated interactions, which can play a critical role in molecular recognition and binding affinity. Halogen bonding, the key interaction exploited by HEF, has been extensively studied since its initial discovery via X-ray crystallography.[3] Its electrostatic nature is now well understood, and it has been shown to significantly influence both selectivity and efficacy in drug-target interactions.[4,5,6]
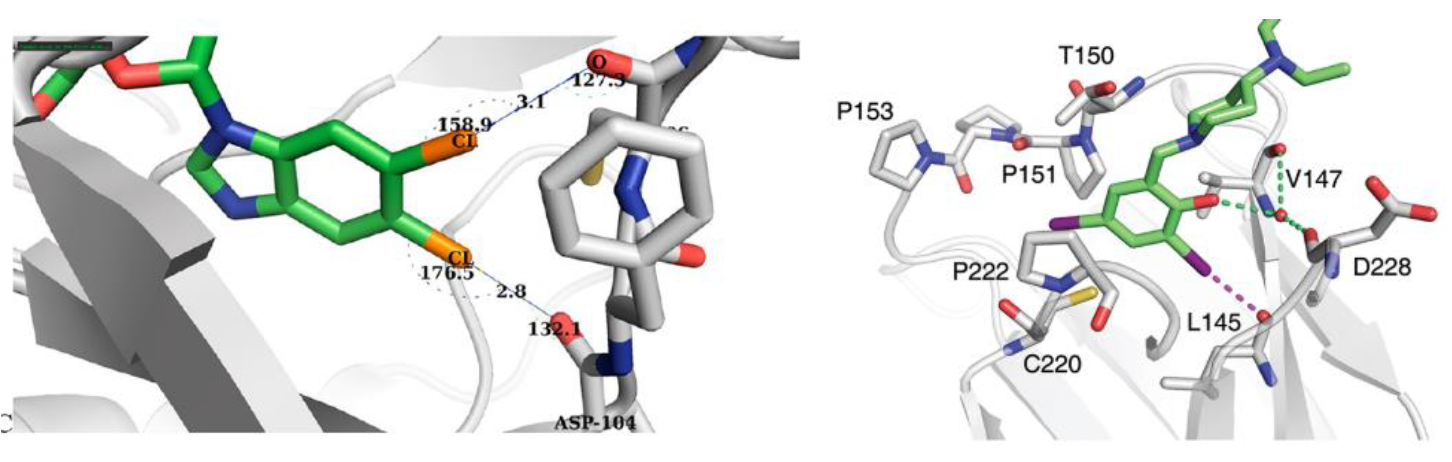
Halogen bonding through σ-hole interactions
A major factor in halogen bonding is the σ-hole, whose characteristics can be “tuned” by varying (hetero)aromatic ring systems and their substitution patterns. This tunability allows for precise control over interaction strength and directionality.
Enamine, a global leader in heterocyclic chemistry, has synthesized thousands of novel halogenated scaffolds, creating the most diverse collection of halogenated core heterocycles available. Our carefully designed Halogen-Enriched Fragment Library comprises the most relevant fragments for exploring affinity towards binding via these non-covalent halogen-mediated interactions.
Typical Formats
Catalog No.
HEF-1920-10-Y-100
Compounds
1 920
6 plates
Amount
10 µL of 100 mM DMSO stock solutions
Plates and formats
384-well microplates, Echo qualified Labcyte, 320 compounds per plate
Price
Catalog No.
HEF-1920-25-X-100
Compounds
1 920
24 plates
Amount
25 µL of 100 mM DMSO stock solutions
Plates and formats
96-well plates, Greiner Cat. No 650201, round (U) bottom, 1 & 12 columns empty, 80 compounds per plate
Price
Catalog No.
HEF-1920-50-X-100
Compounds
1 920
24 plates
Amount
50 µL of 100 mM DMSO stock solutions
Plates and formats
96-well plates, Greiner Cat. No 650201, round (U) bottom, 1 & 12 columns empty, 80 compounds per plate
Price
Key features
- Diverse (hetero)aromatic fragments set for probing halogen bonds
- All compounds include at least one non-fluorine halogen atom
- No PAINS, only MedChem friendly compounds
- Express follow-up from stock and from Enamine REAL
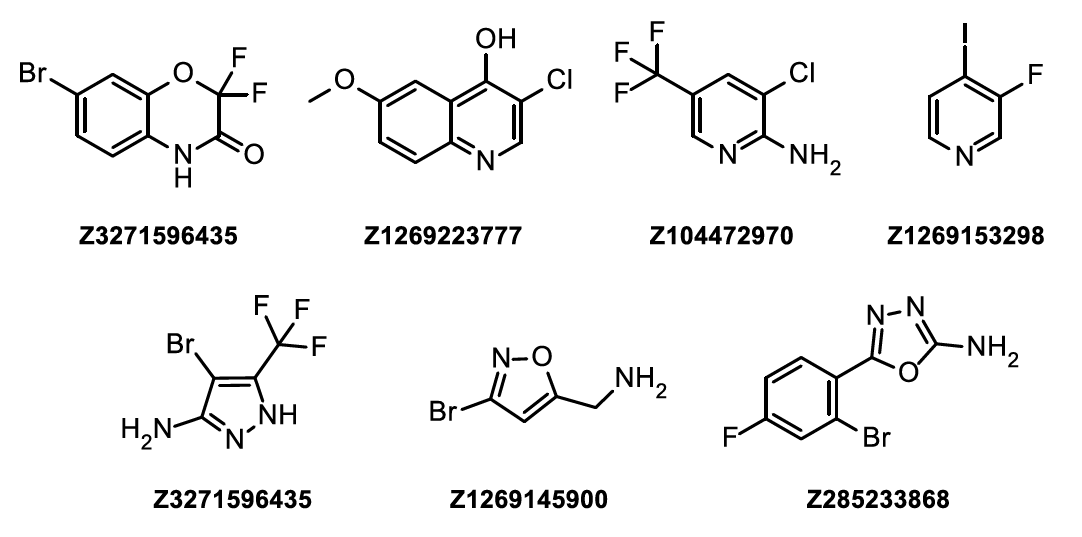
Examples of compounds from HEF Library
Library design
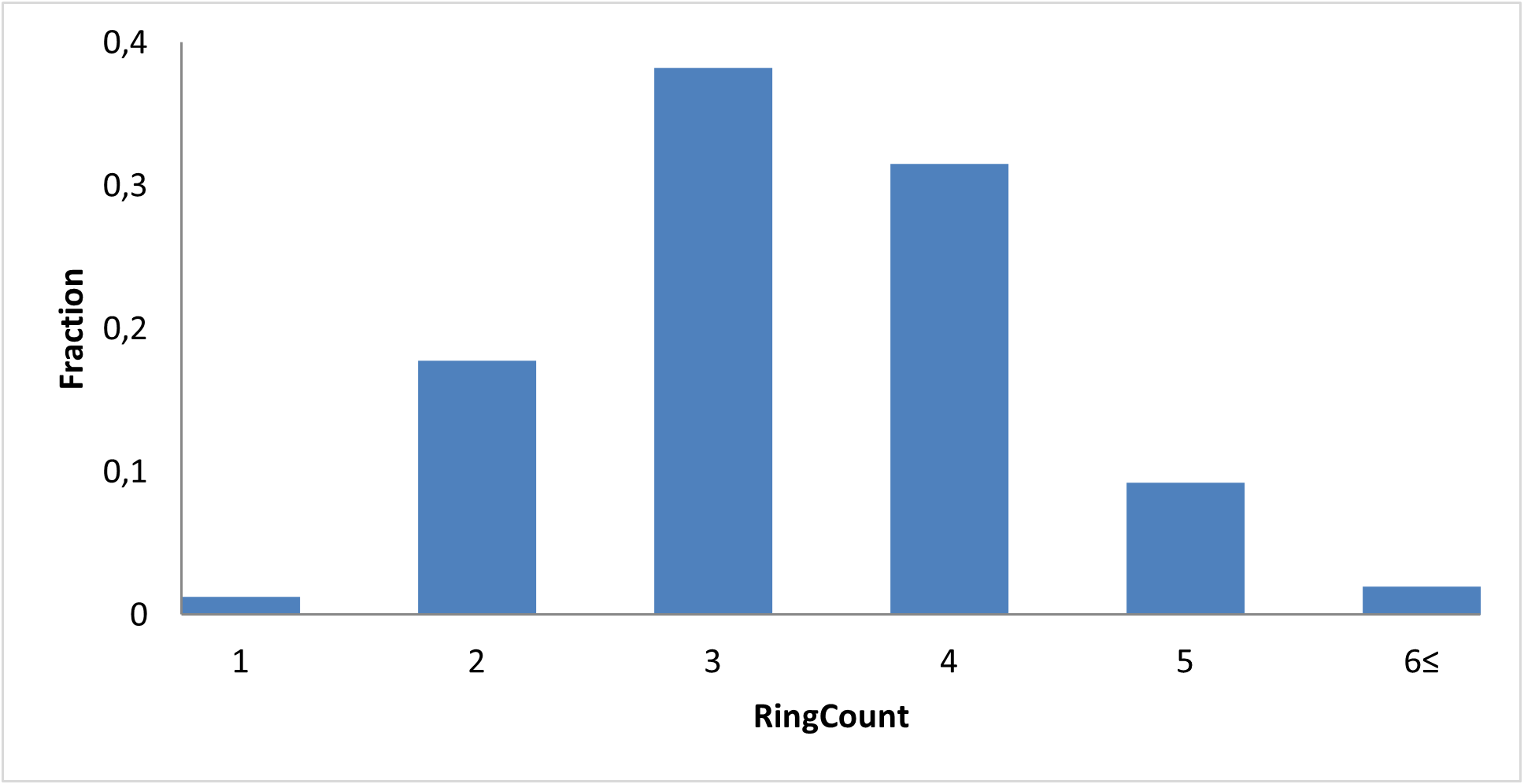
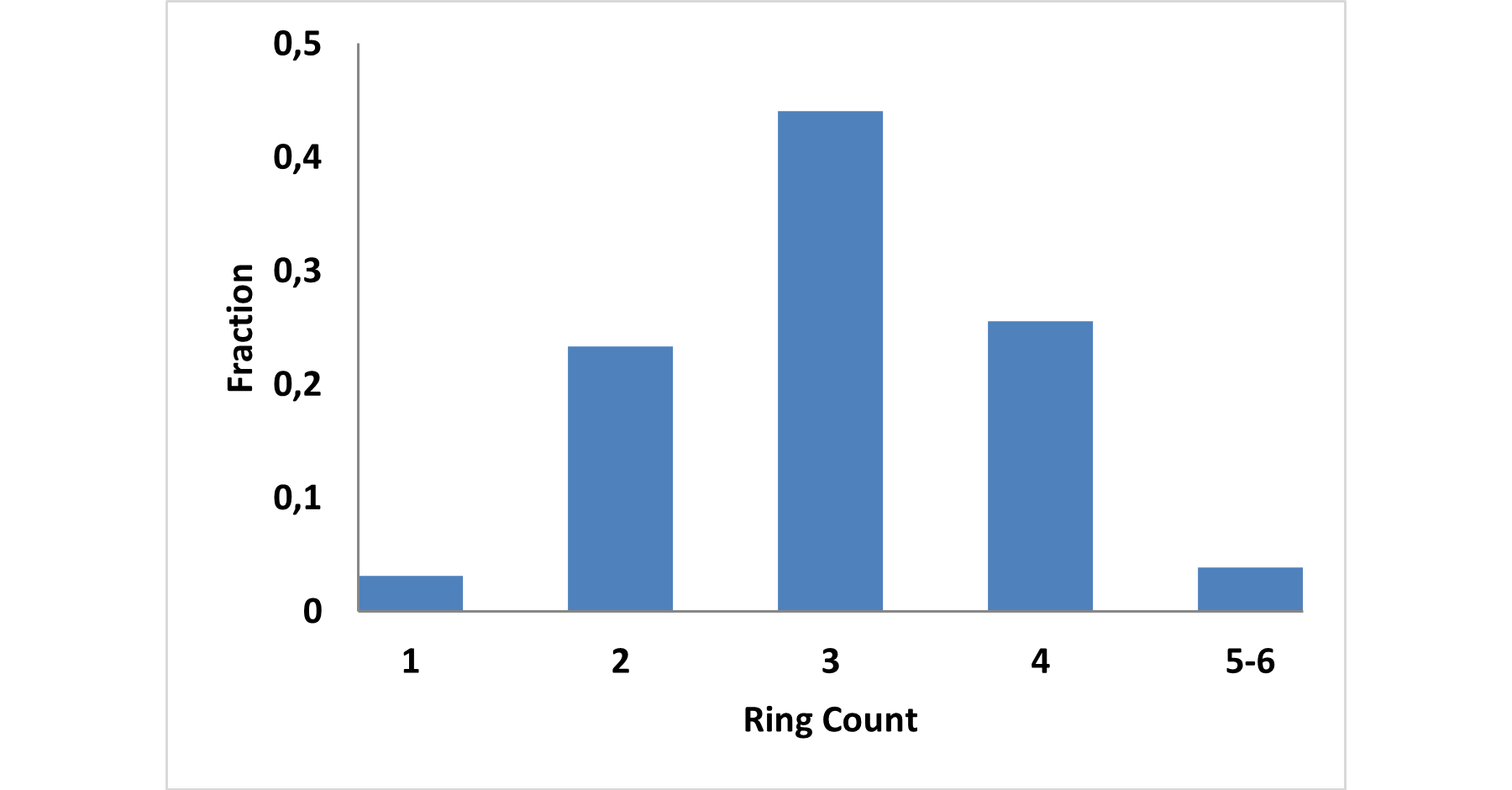
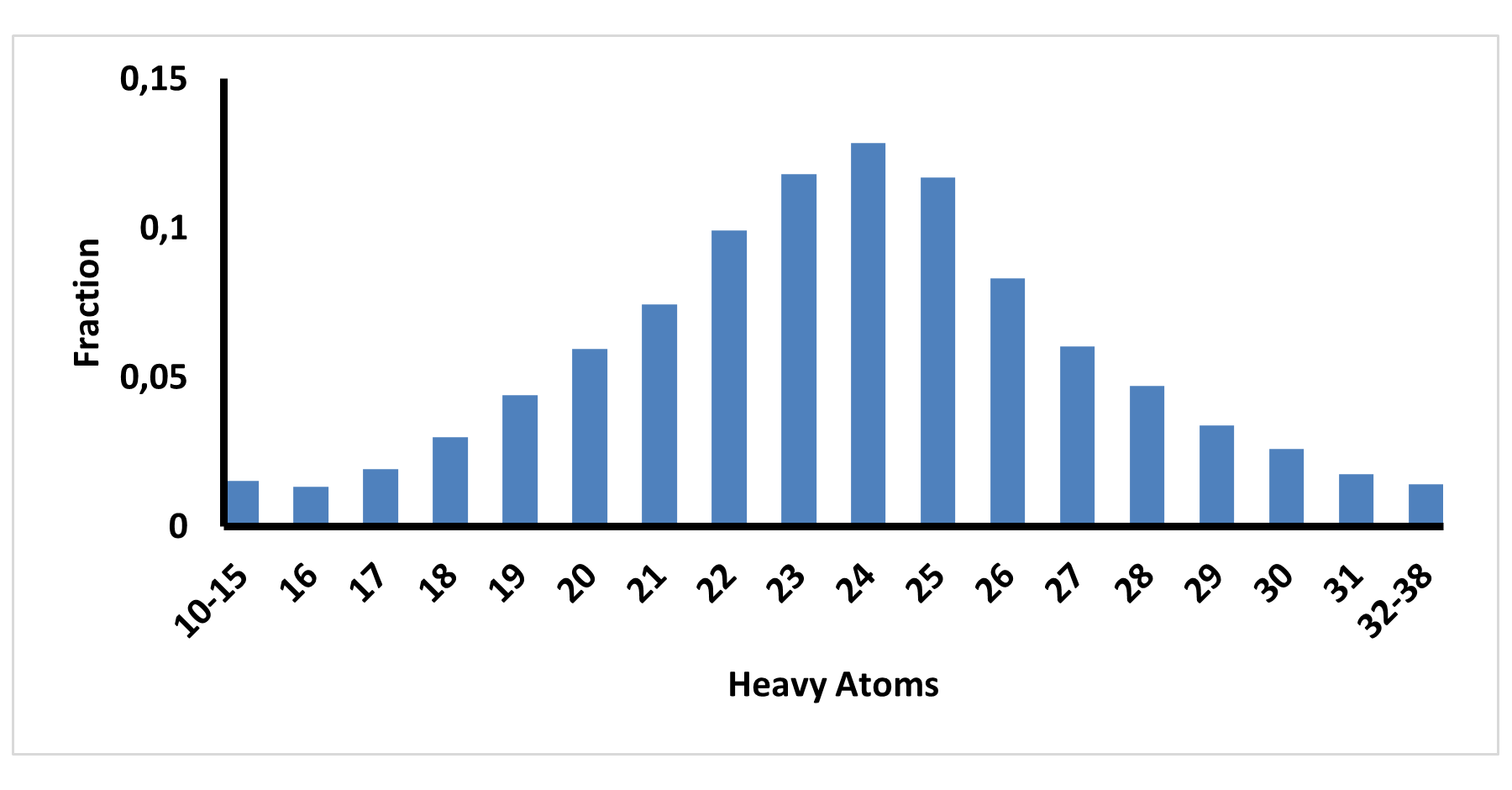
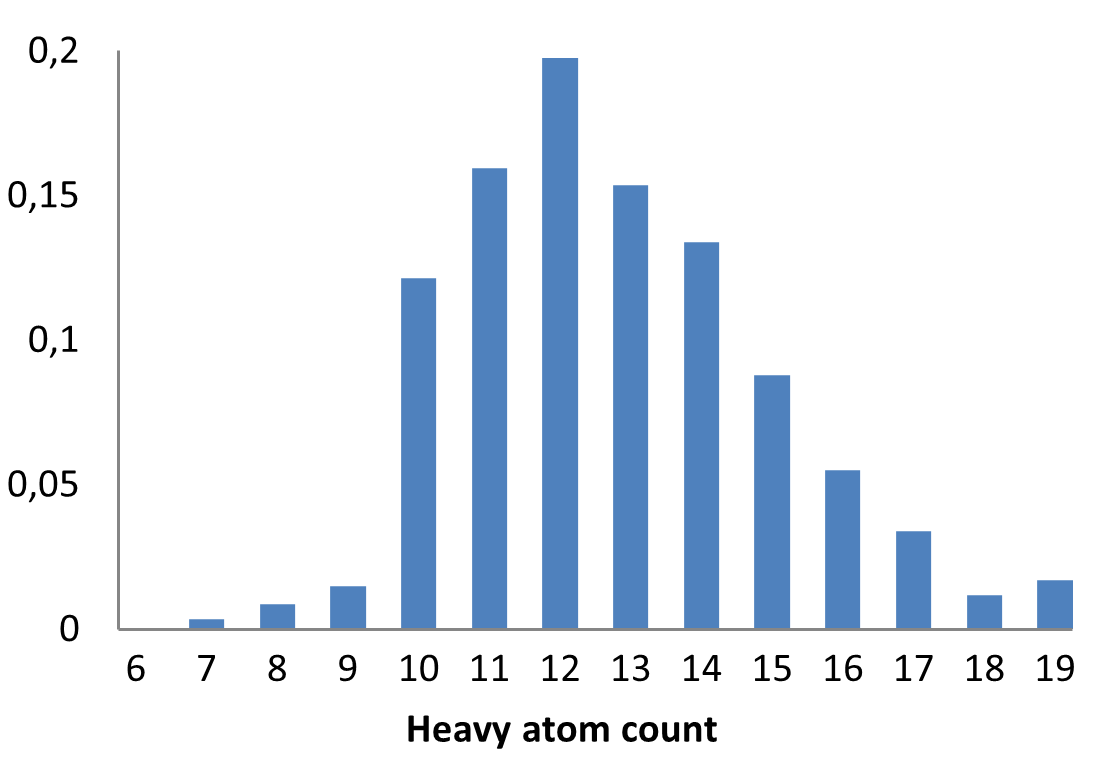
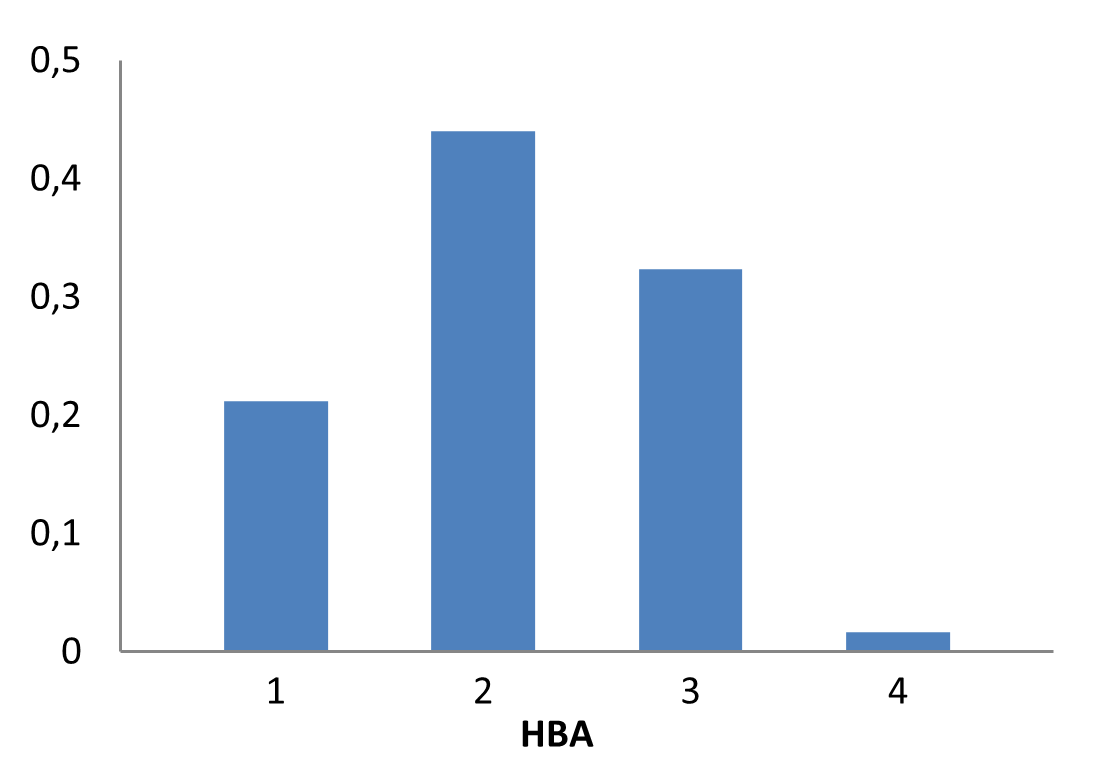
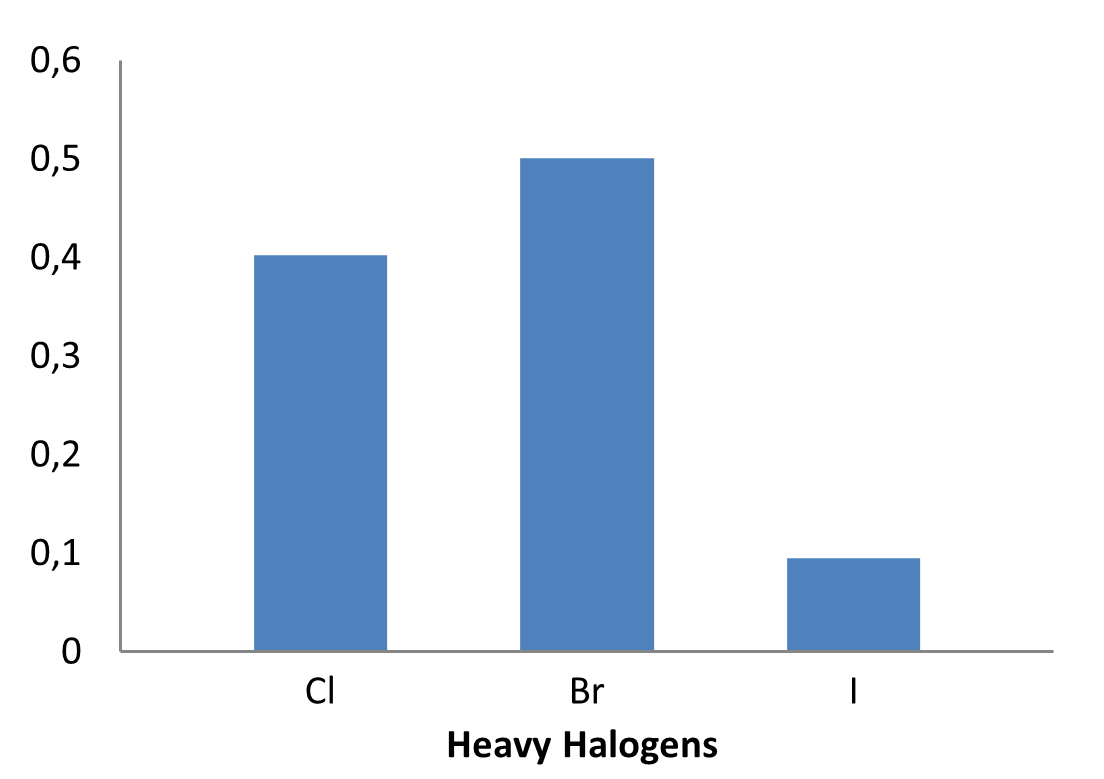
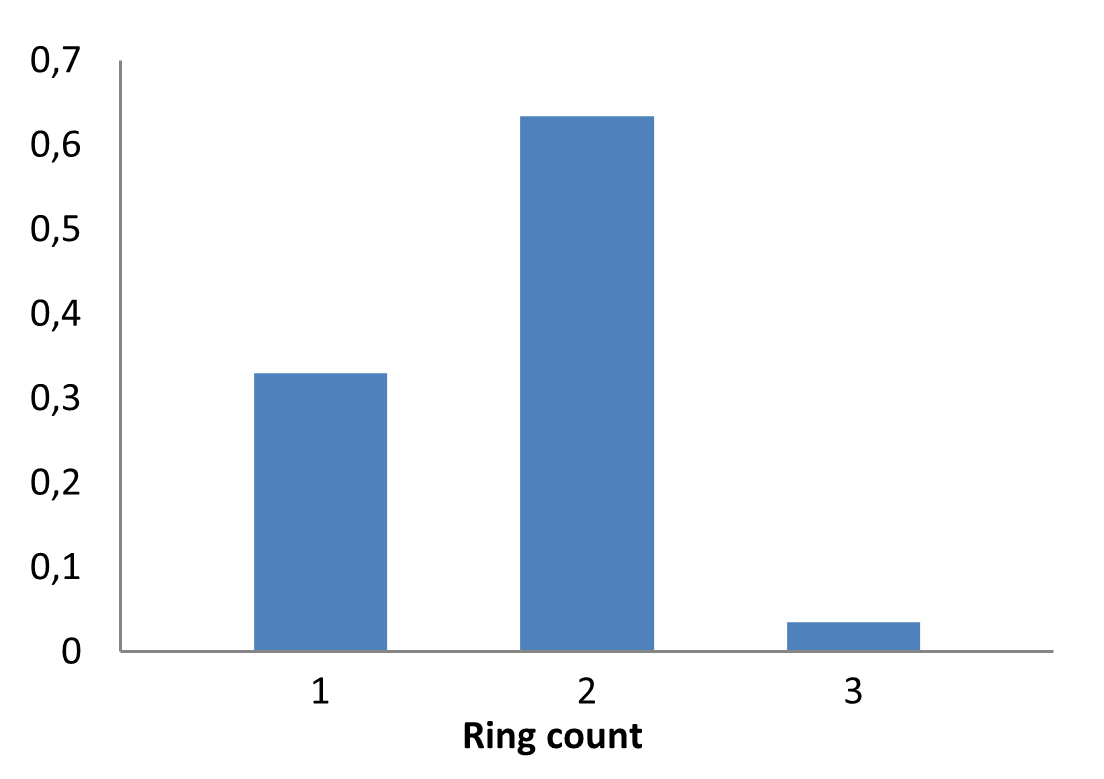
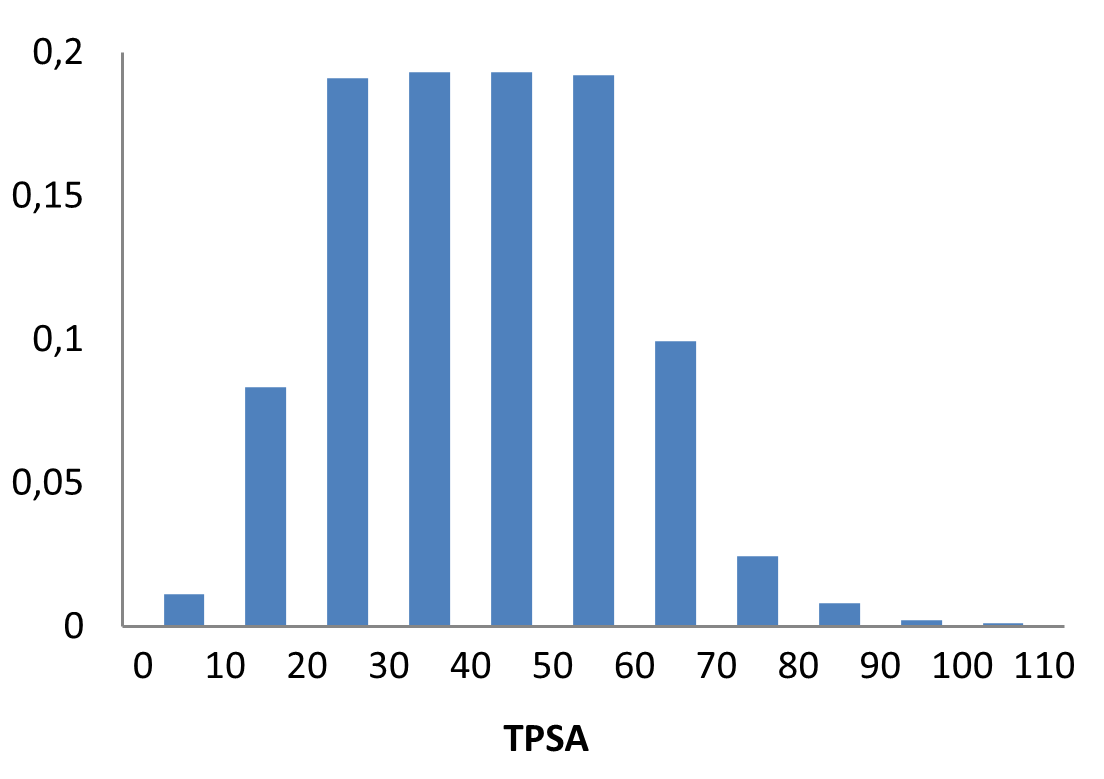
The fragment library was constructed from a curated database of over 250,000 compounds, all strictly compliant with the Rule of Three (Ro3) and filtered using medicinal chemistry criteria. Each compound includes at least one heavy halogen atom (Cl, Br, or I), with 95% containing only one. The distribution is as follows: 183 iodides (9.5%), 963 bromides (50.1%), and 684 chlorides (35.6%). The remaining 4.8% are dichloro derivatives. Additionally, the library includes 194 fluorinated compounds (10.1%). Fragments were selected with heavy atom counts (HA) ranging from 6 to 19, with a majority (1,206 fragments) falling within the 12–16 HA range. Each fragment contains between one and three rings, with 97% featuring one or two. We also included 154 fragments from 200 originally described as HEFLib by Frank M. Boeckler. Scaffold analysis revealed the most frequently occurring (hetero)aromatic ring systems.
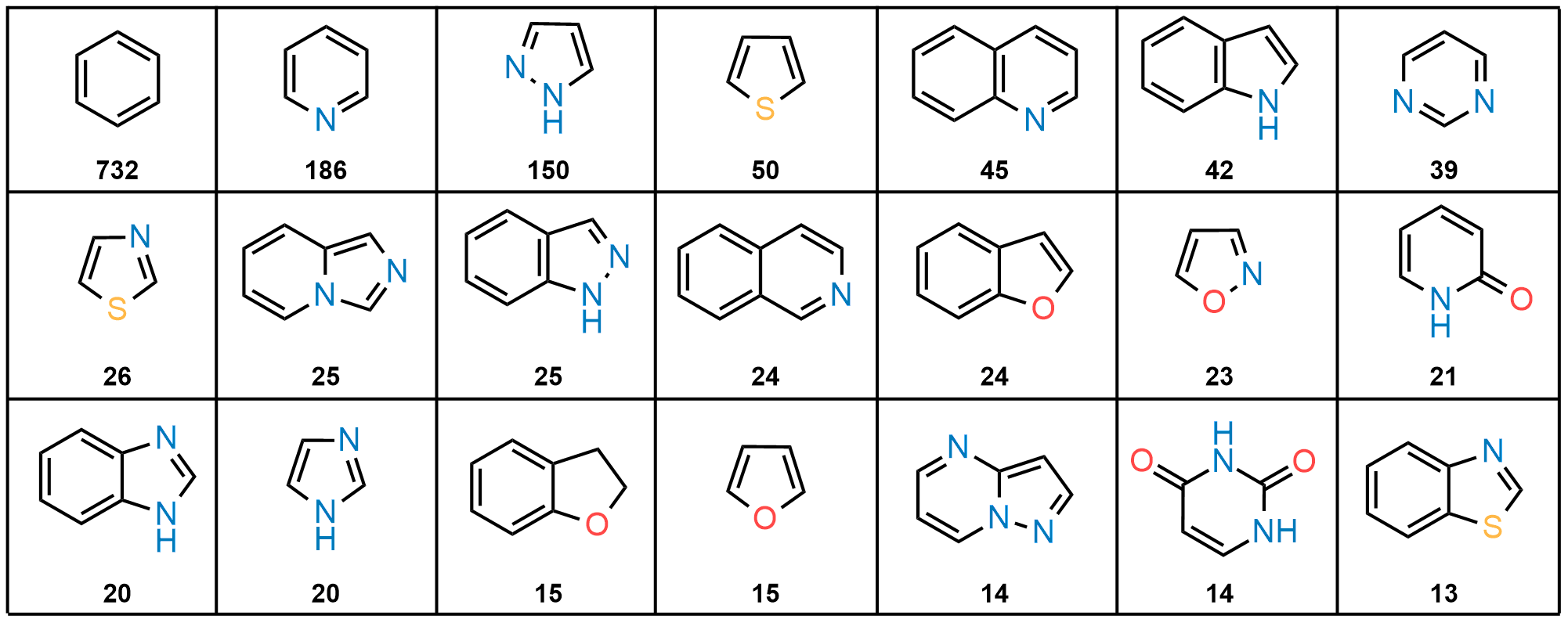
Most Frequently Used Heteroaromatic Ring Systems in the HEF Library
Selected publications
-
Embracing the Diversity of Halogen Bonding Motifs in Fragment-Based Drug Discovery—Construction of a Diversity-Optimized Halogen-Enriched Fragment Library.
J. Heidrich, L. E. Sperl, F. M. Boeckler Front. Chem. 2019, 7. DOI: 10.3389/fchem.2019.00009 -
Halogen interactions in protein-ligand complexes: implications of halogen bonding for rational drug design.
S. Sirimulla, J. B. Bailey, R. Vegesna, M. Narayan J. Chem. Inf. Model. 2013, 53, 2781–2791. DOI: 10.1021/ci400257k -
Ultrahigh resolution drug design I: Details of interactions in human aldose reductase-inhibitor complex at 0.66 Å.
E. I. Howard, R. Sanishvili, R. E. Cachau, A. Mitschler, B. Chevrier, P. Barth, V. Lamour, M. Van Zandt, E. Sibley, C. Bon, D. Moras, T. R. Schneider, A. Joachimiak, A. Podjarny Proteins Struct. Funct. Genet. 2004, 55, 792–804. DOI: 10.1002/prot.20015 -
Halogen bonding: an electrostatically-driven highly directional noncovalent interaction.
P. Politzer, J. S. Murray, T. Clark Phys. Chem. Chem. Phys. 2010, 12, 7748–7757. DOI: 10.1039/C004189K -
Halogen bonding: the sigma-hole.
T. Clark, M. Hennemann, J. S. Murray, P. Politzer J. Mol. Model. 2007, 13, 291–296. DOI: 10.1007/s00894-006-0130-2 -
Halogen-Enriched Fragment Libraries as Leads for Drug Rescue of Mutant p53.
R. Wilcken, X. Liu, M. O. Zimmermann, T. J. Rutherford, A. R. Fersht, A. C. Joerger, F. M. Boeckler J. Am. Chem. Soc. 2012, 134, 6810–6818. DOI: 10.1021/ja301056a
Curated selection of frequent hitters
83 compounds
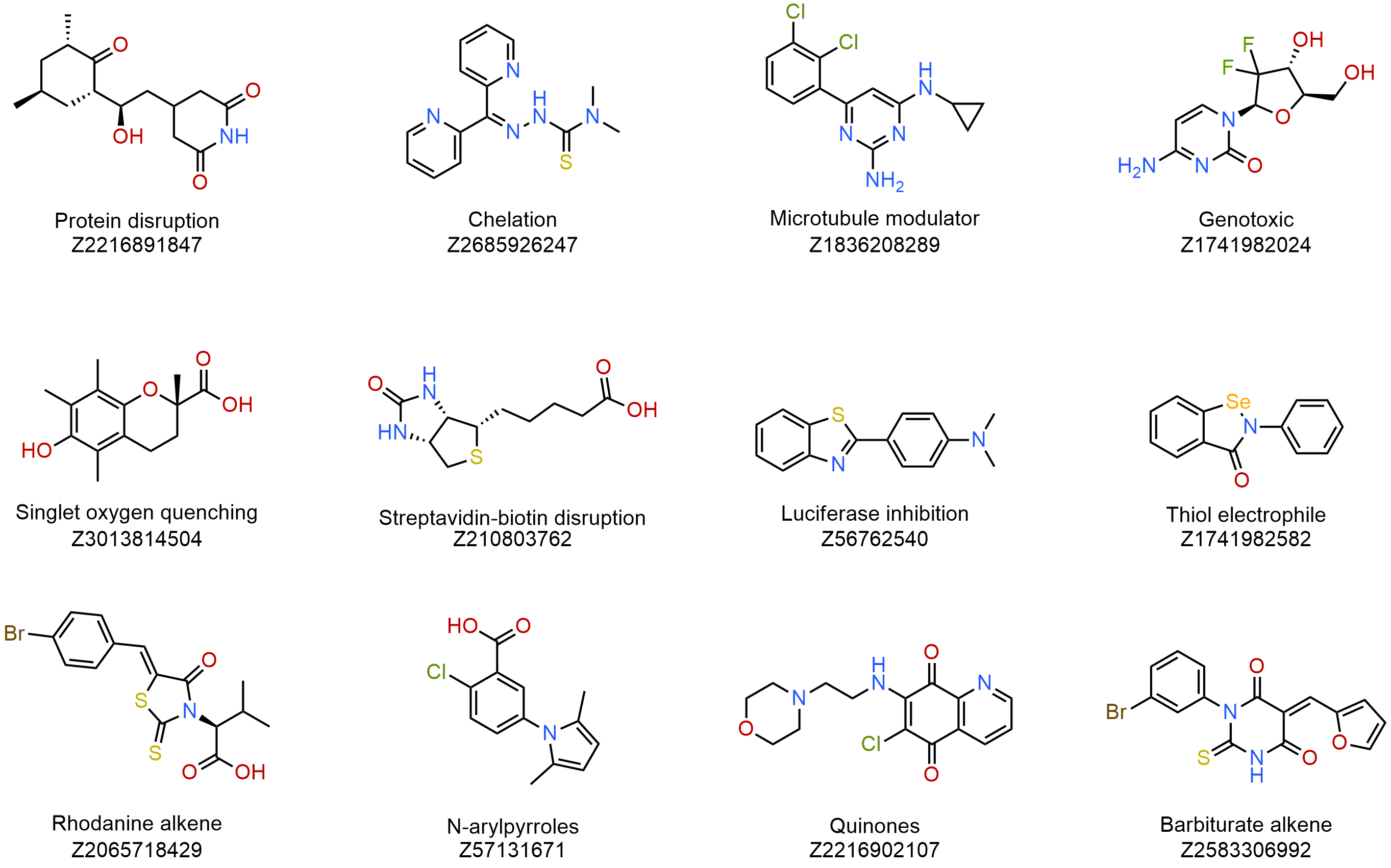
Nuisance compounds are commonly known for their interference with bioassay readouts in high-throughput screening (HTS), causing false positives that are inapplicable to the hit-finding process. Recently, Jonathan Baell provided a description of nuisance compound classes, including colloidal aggregates, PAINS (Pan-Assay Interference Compounds), electrophiles and redox cyclers, chelators, as well as optical and phenotypic assay interference compounds.
We provide a curated list of more than 80 nuisance chemicals known to obstruct assay readouts in target-based and phenotypic screenings. Conveniently available in an assay-ready screening plate, this nuisance compound set is expected to be of great interest to the research community, helping to develop high-quality HTS assays that yield promising, optimizable hits. Hits arising from this small set of nuisance compounds would be highly informative in screening campaigns, assisting target-owners in identifying the types of interference compounds that their assays might be susceptible to.
Most popular library formats available for immediate supply
Catalog No.
CONS-0-Z-10
Compounds
83
1 plate
Amount
≤ 300 nL of 10 mM of DMSO solutions
Plates and formats
1536 well, Echo Qualified 001-6969 (LP-0400), first four and last four columns empty, 1280 compounds per plate
Price
Catalog No.
CONS-10-Y-10
Compounds
83
1 plate
Amount
10 µL of 10 mM DMSO solutions
Plates and formats
384 well, Echo Qualified LDV microplates 001-12782 (LP-0200), first and last two columns empty, 320 compounds per plate
Price
Catalog No.
CONS-50-Y-10
Compounds
83
1 plate
Amount
50 μL of 10 mM DMSO solutions
Plates and formats
384 well, Echo Qualified LDV microplates 001-12782 (LP-0200), first and last two columns empty, 320 compounds per plate
Price
*We will be happy to provide our library in any other most convenient for your project format. Please select among the following our standard microplates: Greiner Bio-One 781270, 784201, 781280, 651201 or Echo Qualified 001-12782 (LP-0200), 001-14555 (PP-0200), 001-6969 (LP-0400) or send your preferred labware. Compounds pooling can be provided upon request.
Download SD file
Key features
- Enhances HTS assay reliability by addressing potential interference
- Over 40 different modes of action
- Readily accessible in an assay-ready screening plate
- References for compounds are available
Library design
The global literature overview and data analysis served as the foundation for the design. Over 80 compounds are categorized based on their specific targets, structural elements, and types of interactions. The collection includes compounds widely recognized and validated as promiscuous across a range of independent and distinct screening methodologies.
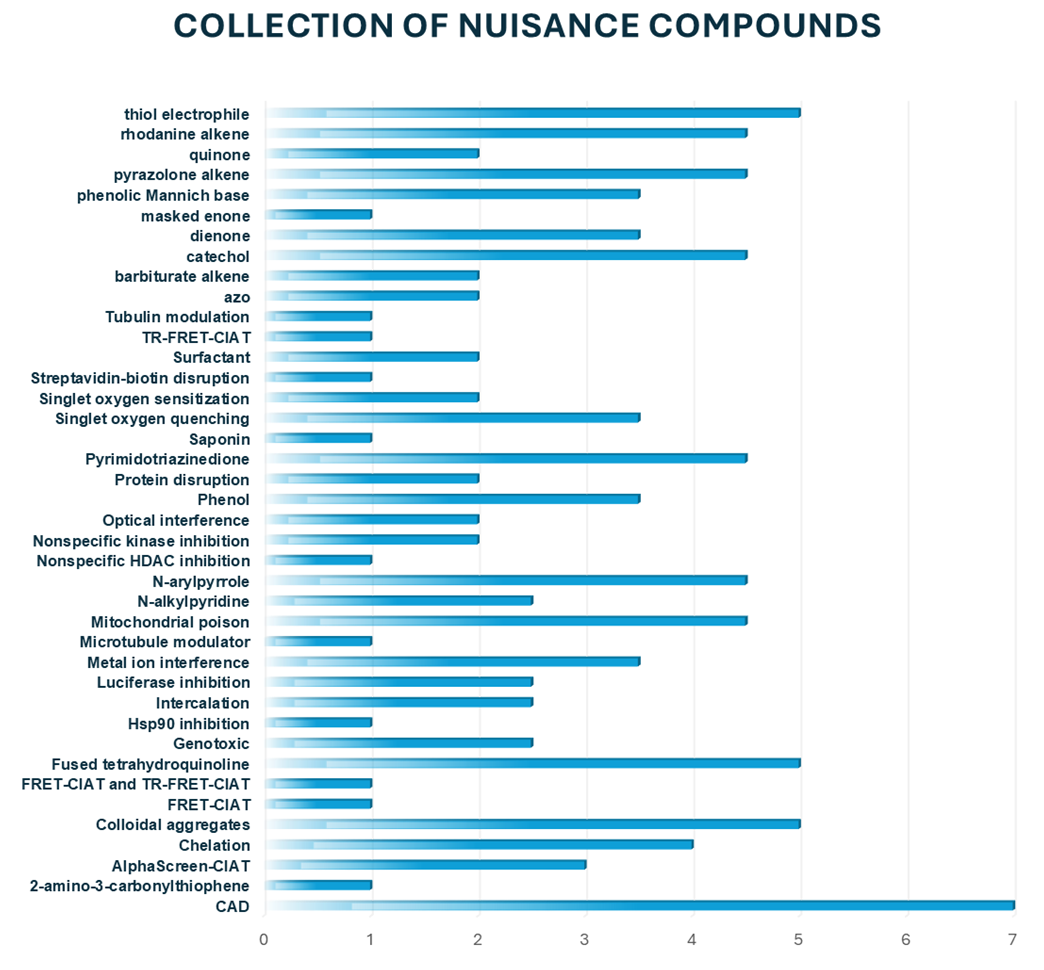
Example of structures