The Synthetically-Accessible Part of Chemical Space for Discovery
Enamine REAL® Compounds are the source and the extension to our in-stock Screening Collection. They are REadily AccessibLe through validated parallel synthesis protocols using our qualified in-stock building blocks, thus representing the synthetically accessible part of chemical space. Ordering the REAL Compounds is as easy as being available from stock.
Some applications of Enamine REAL include:
- Hit exploration
- Hit-to-Lead
- Compound collection enhancement
Quick facts
- Zillions of novel compounds for cherry-picking
- Synthesis within 3-4 weeks, over 80% of compounds delivered on time
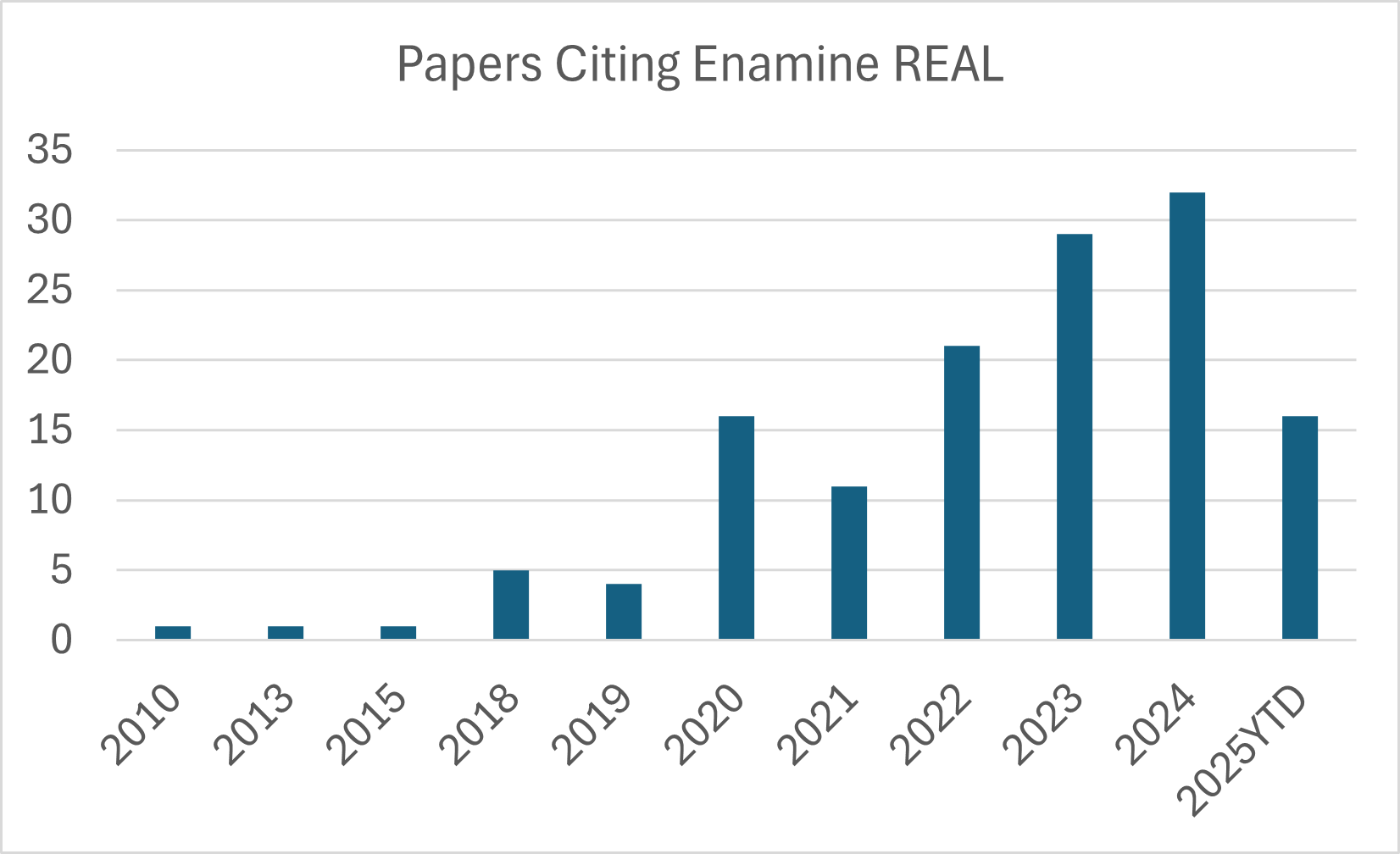
- Recognized by the scientific community, over 130 publications citing Enamine REAL
Explore Enamine REAL:
- Enamine REAL Space. The most recognized and most used subset of Enamine REAL that includes the REAL Database. It can also be accessed via a convenient chemoinformatics tool, infiniSee by BioSolveIT, that generates Enamine REAL Compounds on the fly. Enamine REAL Space is also offered as an enumerated version, provided upon request.
- Enamine REAL Database Subsets: work with the smaller subsets of the REAL Database.
- Enamine xREAL Space. Four! trillion molecules! This subset of Enamine REAL is available through infiniSee xREAL by BioSolveIT and as a part of Chemspace Discovery Services.
- Enamine unREAL Space. A new subset of the REAL compounds that complements the REAL and the xREAL Space. This dataset is provided upon request.
How to get started:
Online search. You can conveniently search for the REAL compounds using substructure and similarity queries on EnamineStore.com, providing access to the entire REAL Database.
Other options:
- Arthor/SmallWorld by NextMove
- Hyperspace plugin for DataWarrior by Alipheron
- ZINC22
- Ignite by Cresset
- Orion (FastROCS) by OpenEye
- KNIME via Chemspace Search Node
- Maestro Suite by Schrödinger
- exaScreen by Pharmacelera
- RIDE by MolSoft
Novelty
Enamine REAL compounds are a next-generation screening compound dataset that allows for exploration of zillions of previously unknown structures. Enamine continuously develops new and innovative building blocks. These building blocks, while combined via well-validated synthetic procedures, provide access to new compounds to drive your projects forward. In most cases, Enamine provides an option of supplying these compounds without publishing them in our stock catalogs and sources like PubChem, Reaxys, and SciFinder.
Diversity
Enamine REAL compounds are based on millions of Murcko scaffolds. We achieved this enormous variety of scaffolds using over 202,223 building blocks assembled via 169 synthetic protocols. Diversity subsets of the REAL compounds are available here.
Drug-likeness
Billions of the REAL compounds are Ro5 and Veber's rule compliant:
- MW ≤500
- SlogP ≤ 5
- HBA ≤ 10
- HBD ≤ 5
- RotBonds ≤ 10
- TPSA ≤ 140
Please use the REAL Database to access these compounds.
Prices
All Enamine REAL Compounds have flat prices. They come from two categories.
- The s-REAL compounds are synthesized via standard 1-2 step procedures with 85%+ success rate.
- The m-REAL compounds are 50% more expensive than the s-REAL compounds because their synthesis involves either multistep synthesis procedures, or use of expensive building blocks, or the products require special purification. Synthesis success rate is over 75%.
- The u-REAL compounds are synthesized via advanced 1-3 step procedures with 65%+ success rate.
Ideas for a quick start
- Diversity set. You can make a pilot screen of REAL compounds using one of the diverse subsets from REAL Database
- Download infiniSee or infiniSee xREAL and run a search within 94.5B or 4 trillion! compounds on an ordinary computer
- Online search. You can conveniently search for the REAL compounds on EnamineStore.com that provides access to the REAL Database.
Yes, Enamine REAL Compounds are not in stock, but...
- All building blocks available in OUR stock
- Each building block has already been tested to comply with the synthesis procedure
- We know in advance which of the synthesis protocols developed in-house to use
...we deliver as if the compounds were available from stock
Selected publications on the use of REAL Compounds
-
Identifying Novel Inhibitors for Hepatic Organic Ani-on Transporting Polypeptides by Machine-learning based Virtual Screening.
Tuerkova A, Bongers B, Norinder U, Ungvári O, Szekely V, Tarnovskiy A, et al. ChemRxiv 2021, in press. DOI: 10.26434/chemrxiv-2021-whpsw -
Structures of the σ2 receptor enable docking for bioactive ligand discovery.
Alon, A., Lyu, J., Braz, J.M. et al. Nature 2021, 600, 759-764. DOI: 10.1038/s41586-021-04175-x -
Synthon-based ligand discovery in virtual libraries of over 11 billion compounds.
Sadybekov, A.A., Sadybekov, A.V., Liu, Y. et al. Nature 2021. DOI: 10.1038/s41586-021-04220-9 -
An open-source drug discovery platform enables ultra-large virtual screens.
Gorgulla, C., Boeszoermenyi, A., Wang, ZF. et al. Nature 2020 580, 663-668. DOI: 10.1038/s41586-020-2117-z -
Design of the Global Health chemical diversity library v2 for screening against infectious diseases.
Wilson, C., Gardner J.M.F., Gray D.W., Baragana B. et al. PLoS Negl Trop Dis. 2023 17(12), e0011799. DOI: 10.1371/journal.pntd.0011799



