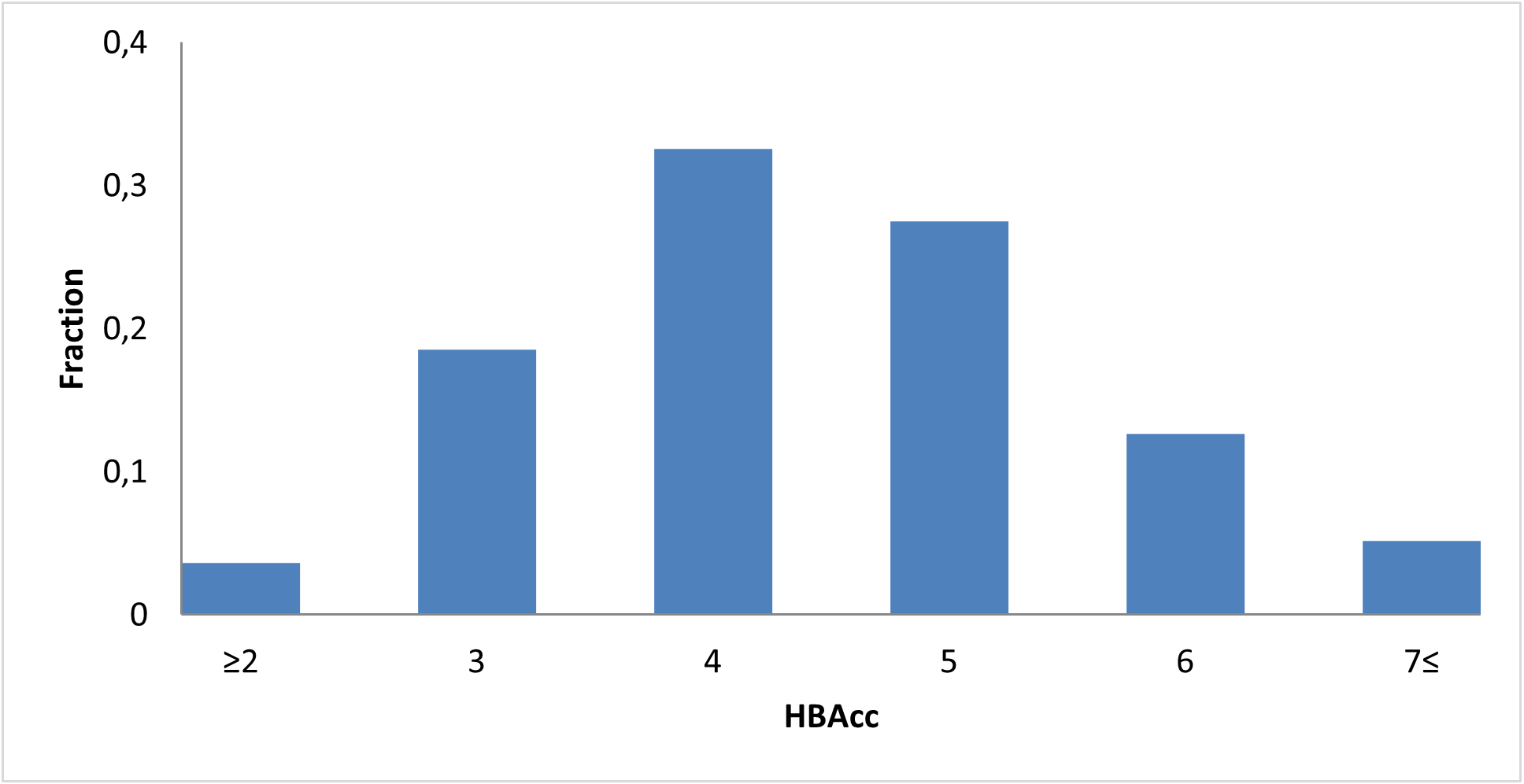Designed to disrupt c-Myc, built to accelerate breakthroughs in cancer
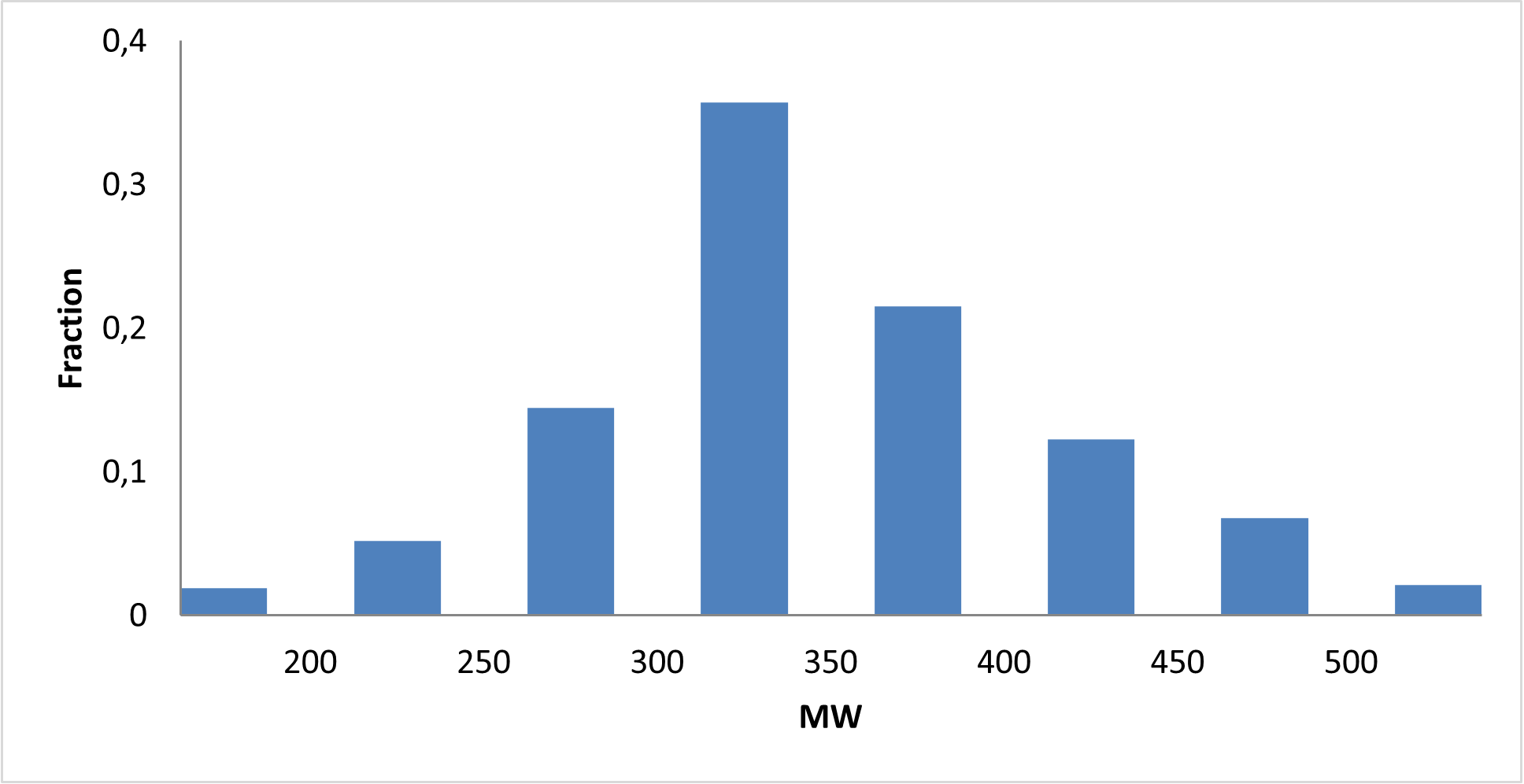
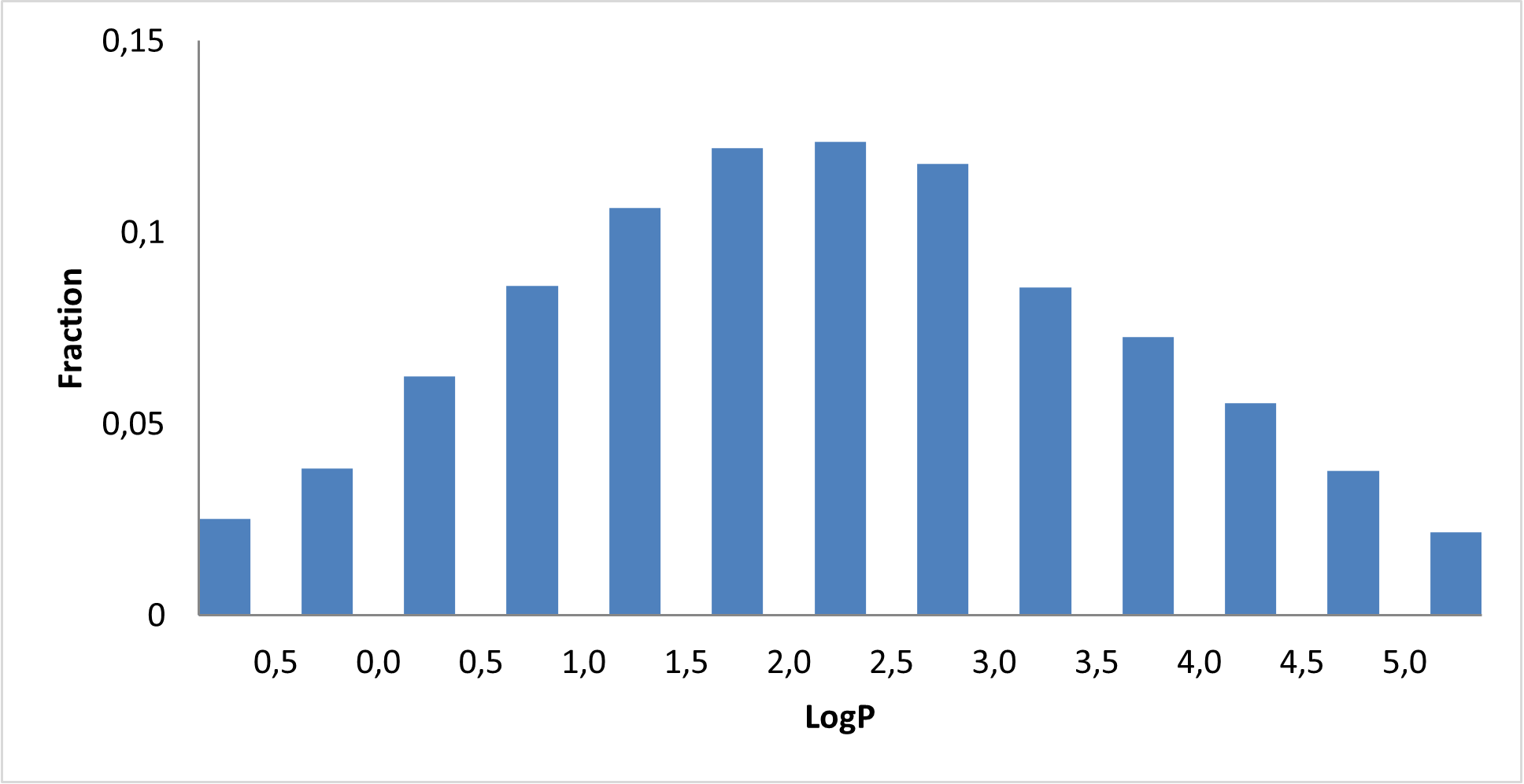
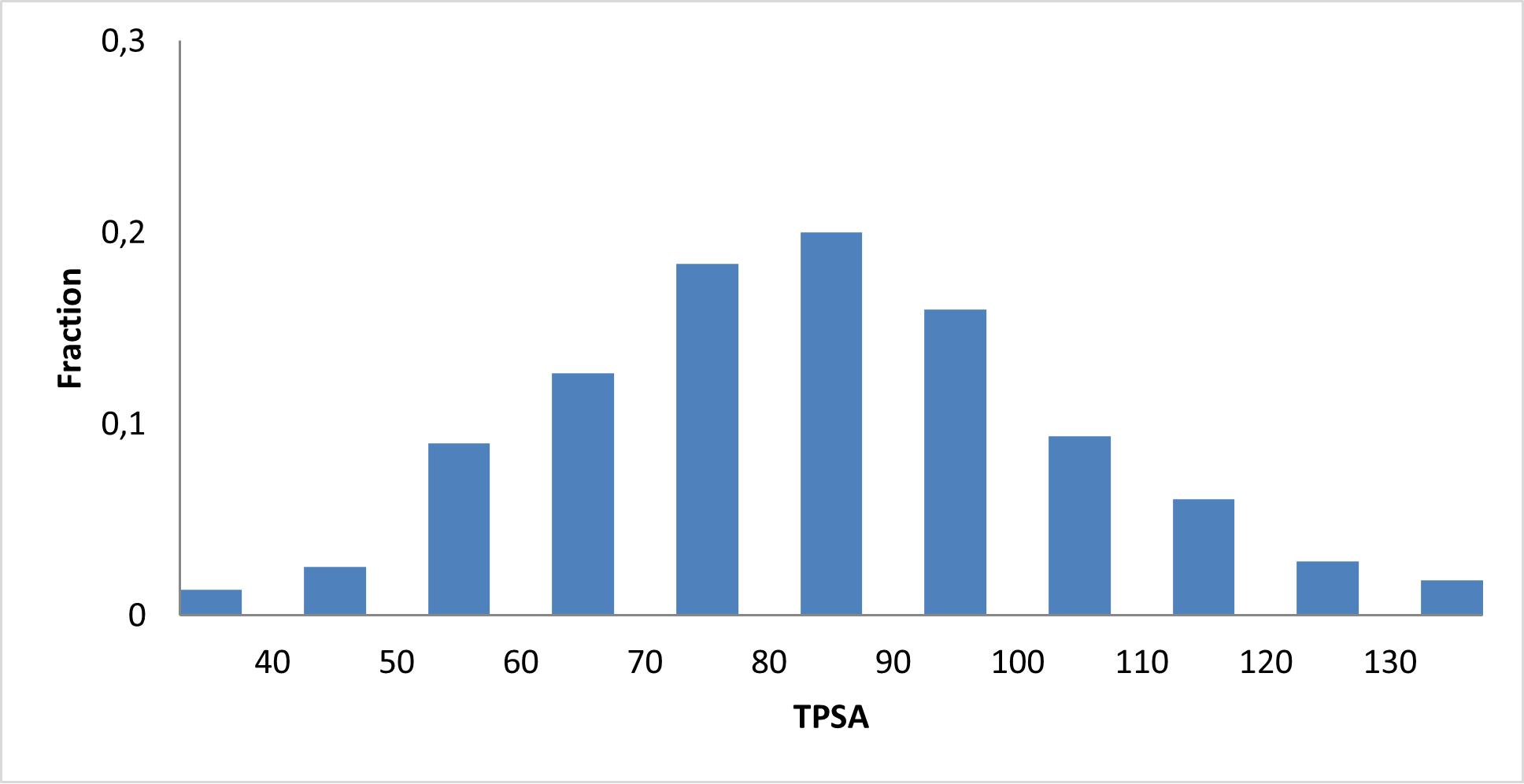
8 160 compounds
The c‑Myc/Max heterodimer is a basic helix–loop–helix leucine zipper (bHLHZip) transcription factor complex that regulates fundamental cellular processes, including growth, differentiation, metabolism, and programmed cell death. Dysregulation of c‑Myc expression is one of the most characteristic oncogenic events in human biology: c‑Myc is constitutively and aberrantly overexpressed in more than half of all human cancers. Its persistent activity promotes uncontrolled proliferation, maintains stem‑cell‑like self‑renewal, and suppresses senescence and differentiation pathways.
Despite its central role in tumorigenesis and decades of research, no clinically approved therapies currently exist that directly target the c‑Myc/Max complex. This gap keeps c‑Myc among the most compelling but challenging targets in modern drug discovery, with relevance spanning oncology, proliferative diseases, and metabolic disorders.
We assembled a curated c‑Myc‑focused library designed to provide high‑quality, mechanism‑relevant starting points for developing novel c‑Myc modulators and chemical probes.
Typical Formats
Catalog No.
MYC-8160-0-Z-10
Compounds
8 160
7 plates
Amount
≤ 300 nL of 10 mM or lower DMSO solutions
Plates and formats
1536 well, Echo Qualified 001-6969 (LP-0400), first four and last four columns empty, 1280 compounds per plate
Price
Catalog No.
MYC-8160-10-Y-10
Compounds
8 160
26 plates
Amount
10 µL of 10 mM DMSO solutions
Plates and formats
384 well, Echo Qualified LDV microplates 001-12782 (LP-0200), first and last two columns empty, 320 compounds per plate
Price
Catalog No.
MYC-8160-50-Y-10
Compounds
8 160
26 plates
Amount
50 μL of 10 mM DMSO solutions
Plates and formats
384 well, Greiner Bio-One microplates 781280, first and last two columns empty, 320 compounds per plate
Price
*We will be happy to provide our library in any other most convenient for your project format. Please select among the following our standard microplates: Greiner Bio-One 781270, 784201, 781280, 651201 or Echo Qualified 001-12782 (LP-0200), 001-14555 (PP-0200), 001-6969 (LP-0400) or send your preferred labware. Compounds pooling can be provided upon request.
Download SD file
Library design
To build a rationally designed compound library against c‑Myc‑related mechanisms, we focused on three complementary strategies:
- Direct Modulation of the c‑Myc/Max Complex
- Targeting the c‑Myc Promoter G‑Quadruplex
- Indirect Inhibition Through Epigenetic and Co‑Regulatory Targets
Although a liganded structure of the c‑Myc/Max complex is still unavailable, several studies have reported small molecules capable of perturbing its function. Leveraging published models and experimentally validated active compounds, we designed two docking models for direct inhibition:
Model 1: Restriction of c‑Myc/Max flexibility near the E‑box binding pocket. Based on the paper J.Med.Chem., we used structural insights surrounding the compound 4DA (JKY‑2‑169), which binds near the c‑Myc/Max–DNA interface and reduces transcriptional activity by limiting conformational dynamics required for DNA recognition.
Model 2: Direct engagement within the E‑box binding interface. Using the concepts reported in Heliyon and Cell Commun Signal, we incorporated the binding mode of XYA1353, a ligand reported to occupy the DNA‑interaction surface of the c‑Myc/Max heterodimer, thereby preventing formation of the transcriptionally active protein–DNA complex.
These two models represent distinct but complementary approaches for identifying modulators that destabilize or prevent c‑Myc/Max engagement with genomic E‑box sequences.
Given the difficulty of drugging intrinsically disordered transcription factors, promoter‑level regulation has emerged as an alternative strategy. A well‑characterized G‑quadruplex structure within the guanine‑rich region of the MYC promoter can repress transcription when sufficiently stabilized. Therefore, small molecules that bind to or stabilize this G‑quadruplex may indirectly suppress c‑Myc activity.
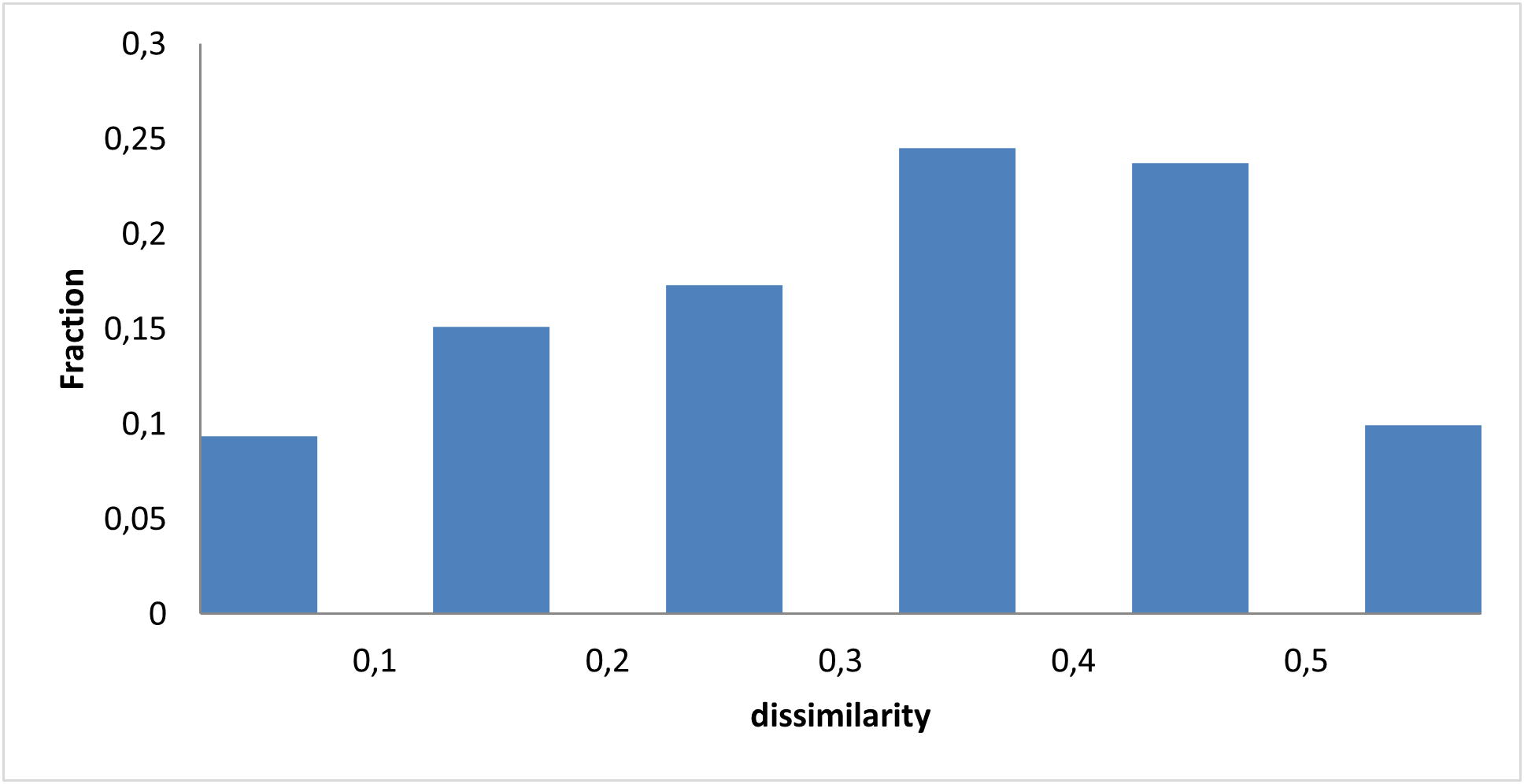
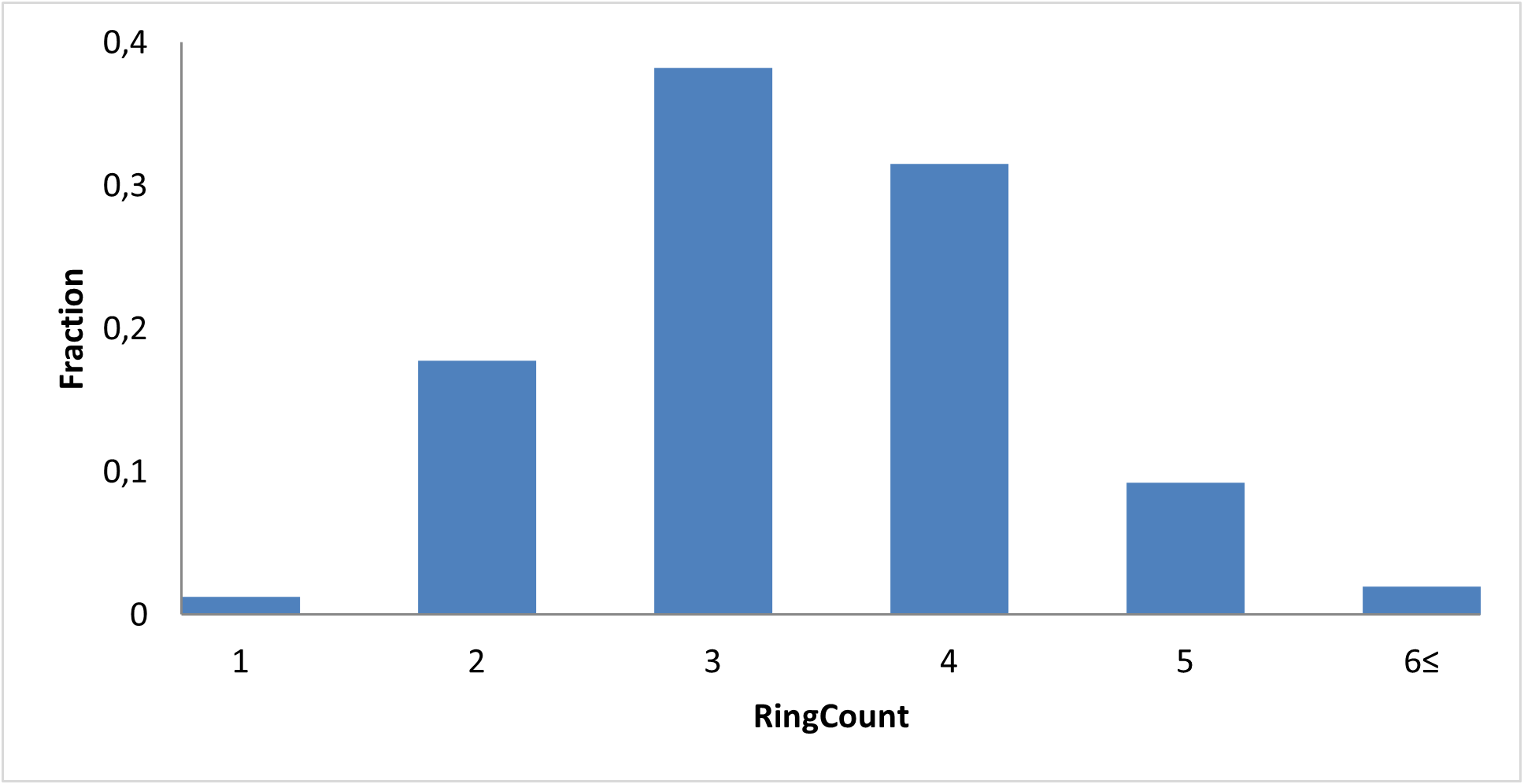
We incorporated major structural features of known modulators: macrocycle–quadruplex complexes, four- and three-fused aromatic rings, and ligands with two linked aromatic rings.
By mapping these structural features, we sought to capture molecules capable of stabilizing the G‑quadruplex and thereby suppressing c‑Myc transcription.
Because c‑Myc/Max relies heavily on chromatin‑associated partners and epigenetic modifiers, indirect inhibition is another promising therapeutic strategy. We focused on targeting key proteins implicated in c‑Myc‑dependent transcriptional regulation: EZH2, HDAC2, KDM4A, KDM4B, KDM4D. These targets are functionally linked to chromatin environments that facilitate c‑Myc–driven gene expression. Modulation of their activity has been shown to alter c‑Myc transcriptional output, making them relevant secondary targets for targeting c‑Myc.
Through these three mechanistic approaches — direct inhibition, promoter G‑quadruplex stabilization, and modulation of c‑Myc‑associated epigenetic regulators — we constructed a diverse and biologically relevant compound library, designed to explore multiple intervention points of c‑Myc regulatory. This multifaceted approach increases the likelihood of identifying promising modulators of the most important oncogenic drivers in human disease.
All protein-ligand complexes were evaluated for stability using a 100 ns MD simulation. The optimized and stable complexes were then used for molecular docking calculations. The following PDB structures have been used for optimization and further simulations – direct binding to c-Myc/Max: 1nkp and 5i50, c-Myc/Max G-quadruplex: 7n7e, 7png, 6jj0, 5w77, and 2a5r, indirect inhibition: 5ls6, 8bpc, 7mos, 7ltk, 7kbh, 6xeb, 6g3o, 8a0b, 7zzw, 7zzt, 5ix0, 3max, 6h4w, 6h4u, 6hgt, 5fpv, 5f3i, 5f3c, 3pdq, 6cg2, 6cg1, 5vmp, 5a7w, 5a7s, 5f5i, 5f39, 3rvh, 6h8p, 7jm5, 5f5c, 5fp4, 5fp8, 6etg, 6ete, 6f5s, 6h0y, and 6h11.
MD simulation result
Model 1
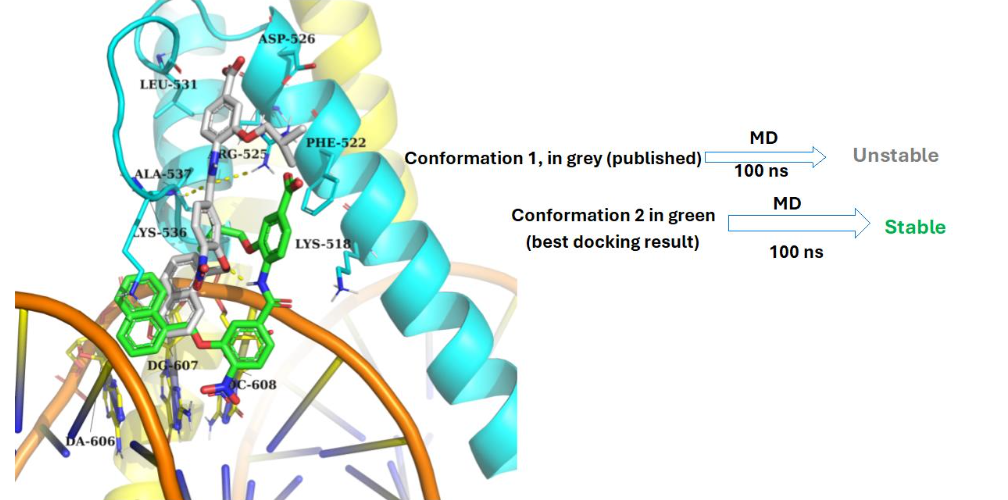
Binding models between c-Myc/Max and 4DA (“JKY-2-169”). DNA in orange, Myc in yellow, Max in cyan.
Model 2
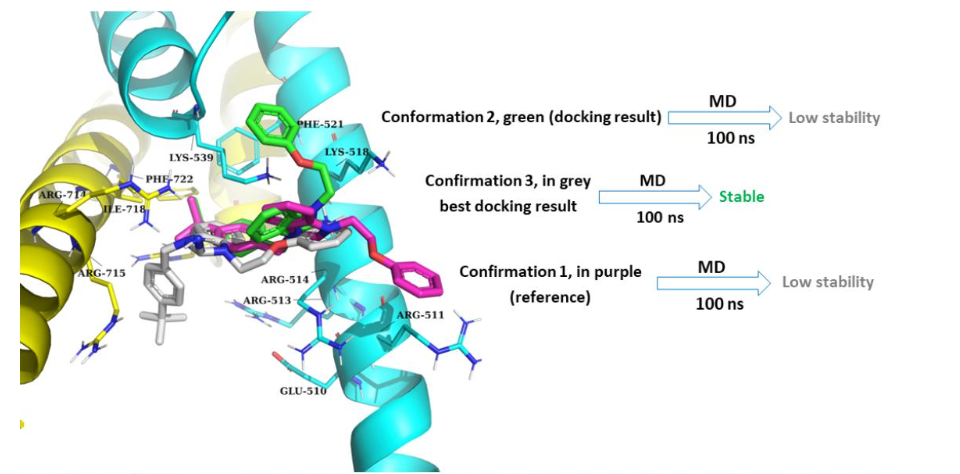
Binding models between c-Myc/Max and XYA1353. DNA in orange, Myc in yellow, Max in cyan.
Examples of molecular docking simulation, and used pharmacophore models, c-Myc/Max protein, direct interaction
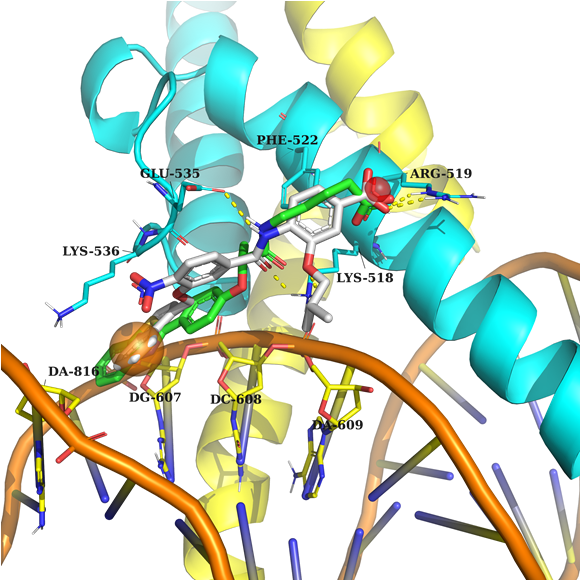
Example 1 (PDB ID 1nkp, c-Myc/Max)
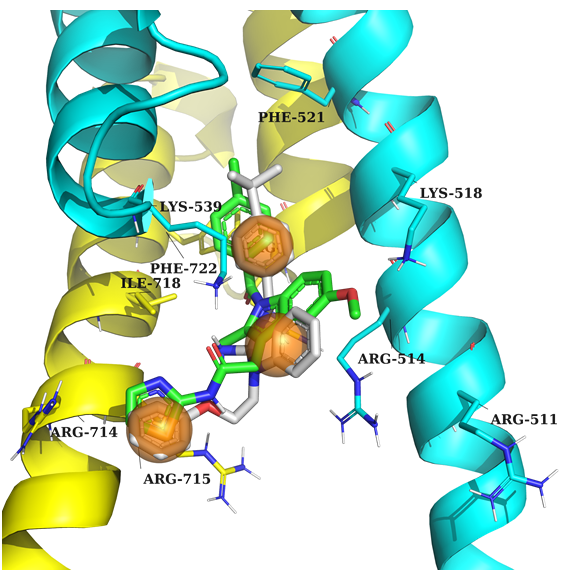
Example 2 (PDB ID 1nkp, c-Myc/Max)
Example 1: According to the pharmacophore model, the ligand's acceptor interacts with Arg 519, and the aromatic ring fills the sub-pocket created by Lys 536 and the nucleotide environment of DNA.
Example 2: Key feature – ligand should contain 3 aromatic rings to create stacking interaction with nearby amino acids.
DNA is in orange, Myc in yellow, Max in cyan, native ligand in grey, docked example in green.
Examples of molecular docking simulation, and used pharmacophore models, c-Myc/Max G-quadruplexes
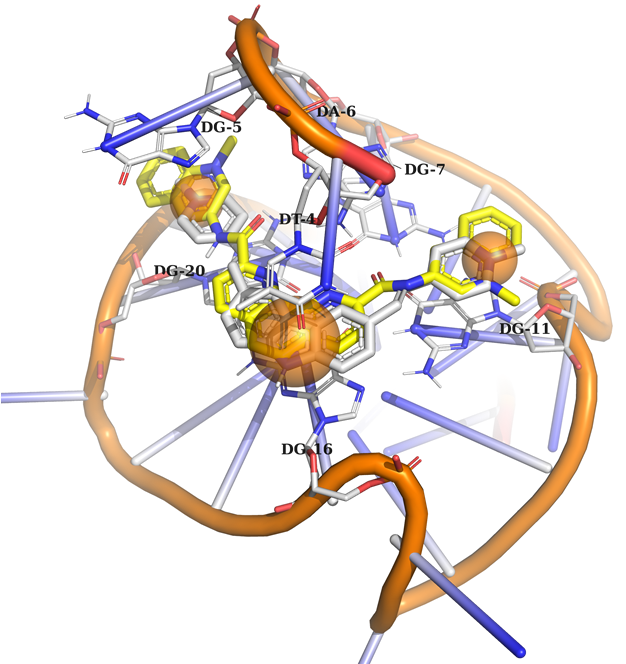
Example 1 (PDB ID 6jj0, c-MYC G-quadruplex)
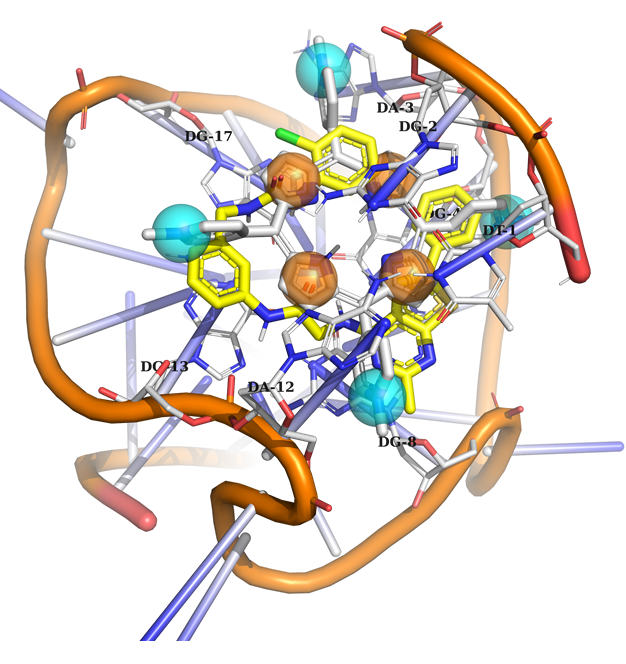
Example 2 (PDB ID 2a5r, c-MYC G-quadruplex)
Example 1: Ligand should contain 3 aromatic rings to create a stacking interaction with the nucleotides.
Example 2: The model contains 8 binding points, including 4 aromatic and mixed aromatic or N+ feature. According to this model, a potential ligand must occupy at least 3 out of 8 binding points to be selected.
DNA in orange, native ligand in grey, docked ligand in yellow.
Examples of molecular docking simulation, and used pharmacophore models, Indirect inhibition strategies
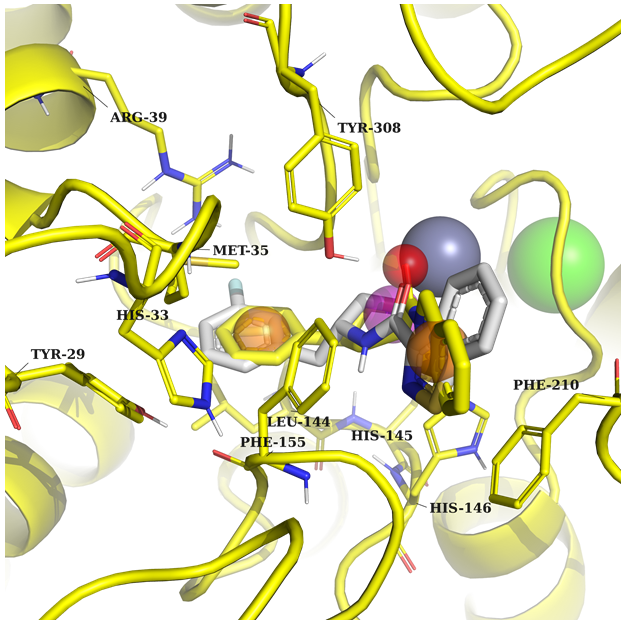
Example 1 (PDB ID 8a0b, HDAC2)
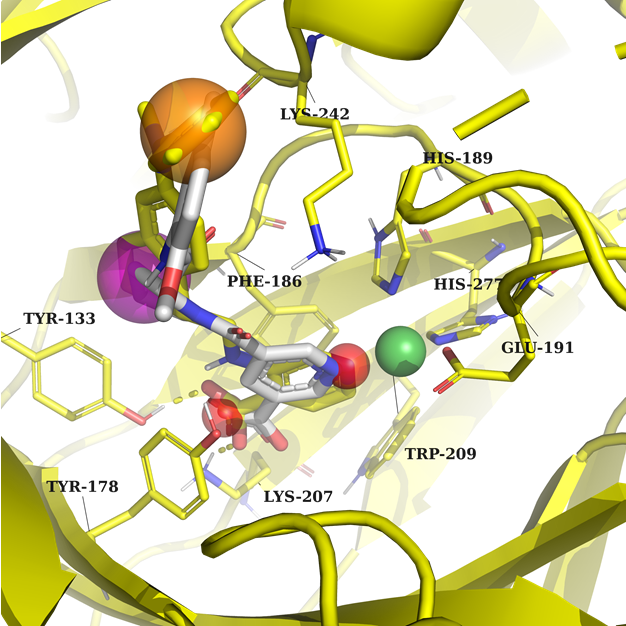
Example 2 (PDB ID 7jm5, KDM4B)
Example 1: A potential ligand requires two aromatics to create π-stacking interaction with Phe155, Phe210, and Tyr308 in one part, and Arg39 and Met35 on the other part of the pocket. Key features: acceptor, ring and donor near the metal atom.
Example 2: Two acceptor features interacting with Lys207, Tyr133, and a metal. Any ring or aromatic ring to fill the binding pockets and create interaction with Lys242.
In both cases, the protein and native ligand are in yellow, and the docked structure in grey.