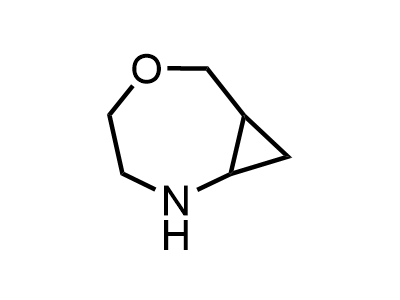
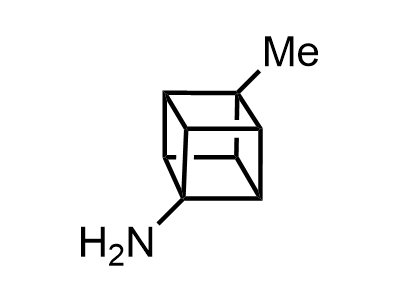
300 Thousand compounds in stock
Original and unique
Make-on-demand
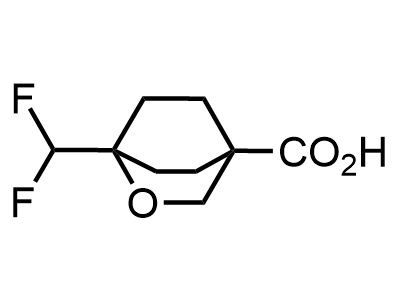
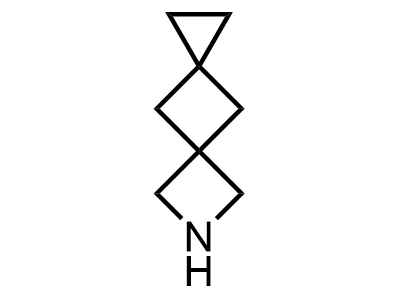
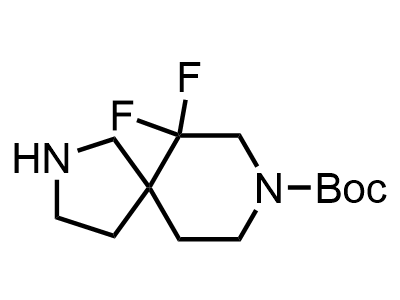
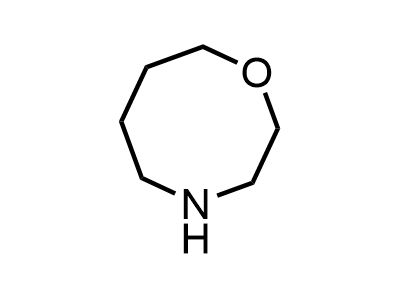
Building Blocks
1B novel building blocks
Reliable supply
Over 650 highly skillful chemists
Unique synthesis technologies
48B Billion
REAL compounds and
Custom Library Synthesis
On site access to all Enamine stock BB’s
Highly flexible arrangements
2 000 new building blocks are synthesized monthly. Here is an important update to our MedChem Highlights from March 2024
Recent News
11 April 2024
Press Release
Cambridge, UK and Kyiv, Ukraine, 11 April 2024: Metrion Biosciences Limited (“Metrion”), the specialist ion channel and cardiac safety screening contract research organisation (CRO) and drug discovery company, and Enamine Ltd (“Enamine”), the global leader in supplying small molecules and early drug discovery services, announced that Metrion has enhanced its High Throughput Screening (HTS) services with the addition of access to Enamine’s compound libraries.
27 March 2024
Press Release
March, 2024, Kyiv, Ukraine. Enamine Ltd, the global leader in supplying small molecules and early drug discovery services, announces the expansion of its library synthesis capabilities with a focus on Enamine REAL compounds to further support the growing demands of agricultural and pharmaceutical companies, research institutes, and drug discovery centers.
01 March 2024
News
We are excited to announce a strategic collaboration between Enamine, the world's leading provider of chemical building blocks, compound libraries, and biology services, and Genez International, a prominent enterprise with 15 years of experience in cross-border supply management, biopharmaceutical research and development, semiconductor equipment, and high-definition digital imaging systems.
ACS Omega
2021, 6 (6), 4227-4235
DOI:
10.1021/acsomega.0c05143
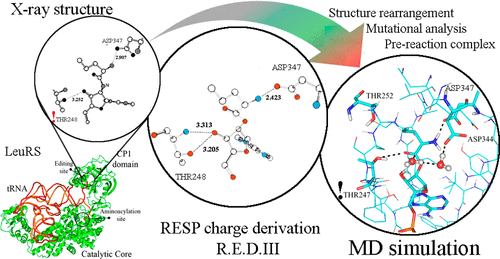
Rayevsky A.; Sharifi M.; Demianenko E.; Volochnyuk D.; Tukalo M.
An important aspect of molecular mechanics simulations of a protein structure and ligand binding often involves the generation of reliable force fields for nonstandard residues and ligands. We consider the aminoacyl-tRNA synthetase (AaRS) system that involves nucleic acid and amino acid derivatives, obtaining force field atomic charges using the restrained electrostatic potential (RESP) approach. These charges are shown to predict observed properties of the post-transfer editing reaction in this system, in contrast to simulations performed using approximate charges conceived based upon standard charges for related systems present in force field databases. In particular, the simulations predicted key properties induced by mutation. The approach taken for generating the RESP charges retains established charges for known fragments, defining new charges only for the novel chemical features present in the modified residues. This approach is of general relevance for the design of force fields for pharmacological applications, and indeed the AaRS target system is itself relevant to antibiotics development.