Library of compounds intended for use in agro/crop science
15 084 compounds
Typical Formats
Catalog No.
AGR-10-0-Z-10
Compounds
10 240
8 plates
Amount
≤ 300 nL of 10 mM of DMSO solutions
Plates and formats
1536-well Echo LDV microplates, first and last four columns empty, 1280 compounds per plate
Price
Catalog No.
AGR-10-10-Y-10
Compounds
10 240
32 plates
Amount
≤ 10 µL of 10 mM DMSO solutions
Plates and formats
384-well, Echo Qualified LDV microplates #001-12782 (LP-0200), first and last two columns empty, 320 compounds per plate
Price
Catalog No.
AGR-10-50-Y-10
Compounds
10 240
32 plates
Amount
50 μL of 10 mM DMSO solutions
Plates and formats
384-well, Greiner Bio-One plates #781280, 1,2 and 23,24 columns empty, 320 compounds per plate
Price
Catalog No.
Library & follow-up package
Plates and formats
AGR-10-10-Y-10 screening library 10 240 cmpds, hit resupply, analogs from 4.7M+ stock and synthesis from REAL Space
Price
*We will be happy to provide our library in any other most convenient for your project format. Please select among the following our standard microplates: Greiner Bio-One 781270, 784201, 781280, 651201 or Echo Qualified 001-12782 (LP-0200), 001-14555 (PP-0200), 001-6969 (LP-0400), C52621 or send your preferred labware. Compounds pooling can be provided upon request.
Download SD files
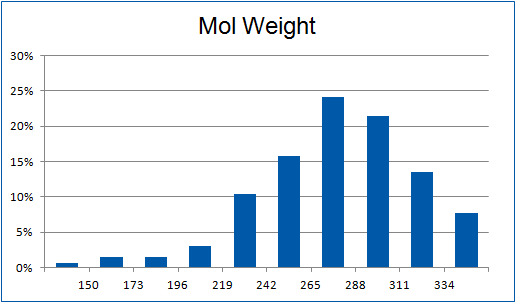
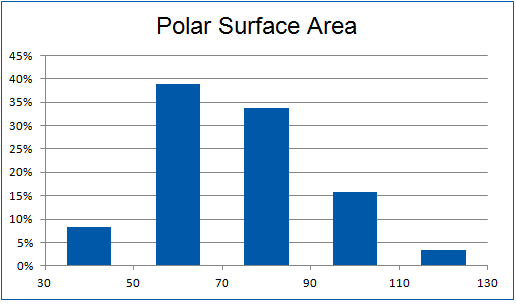
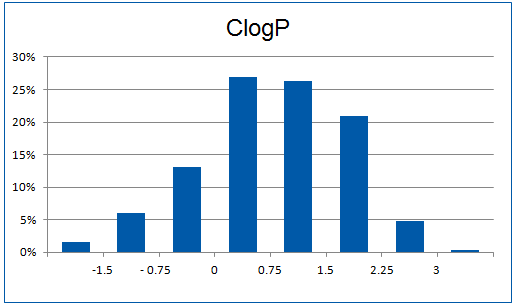
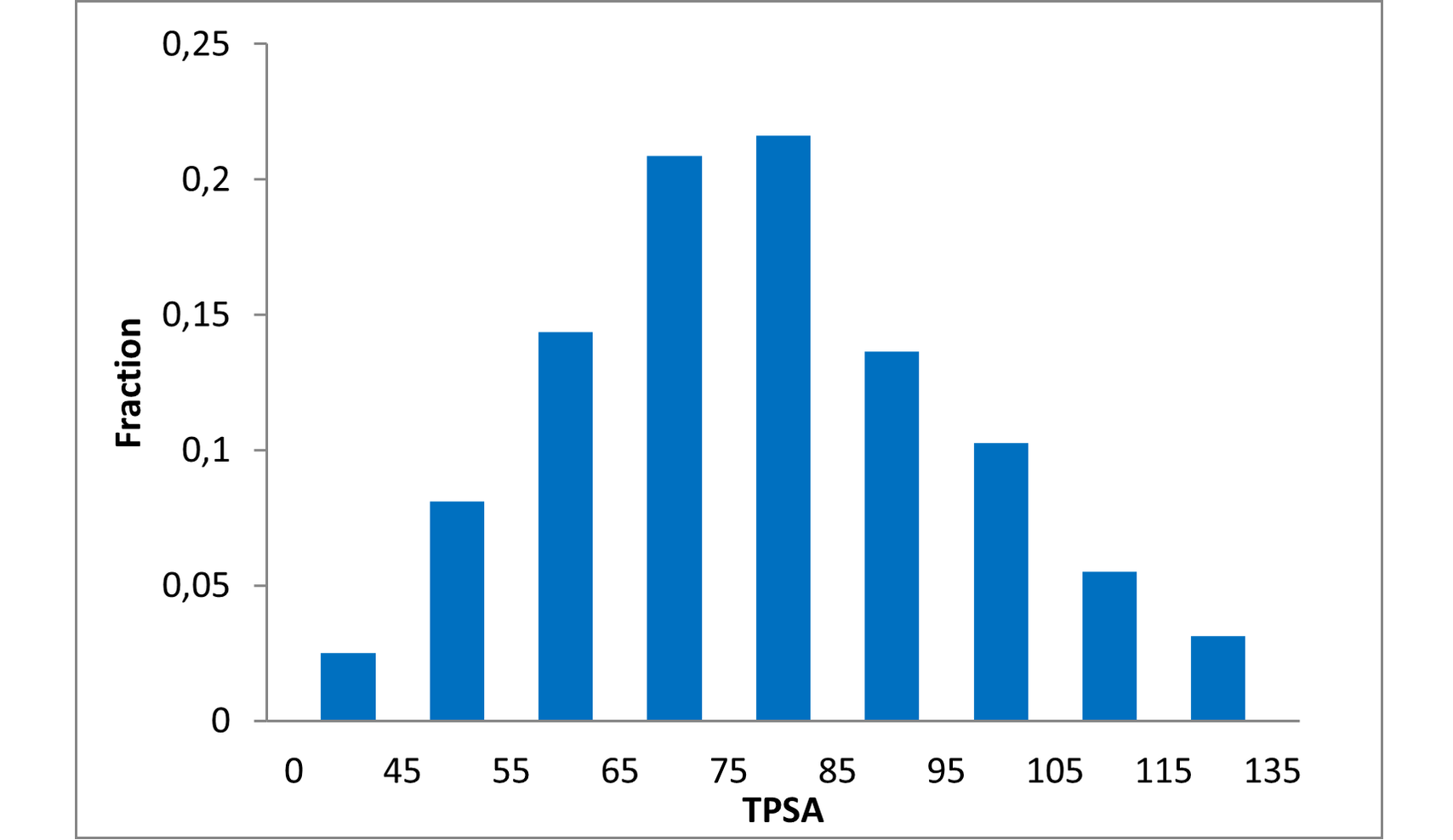
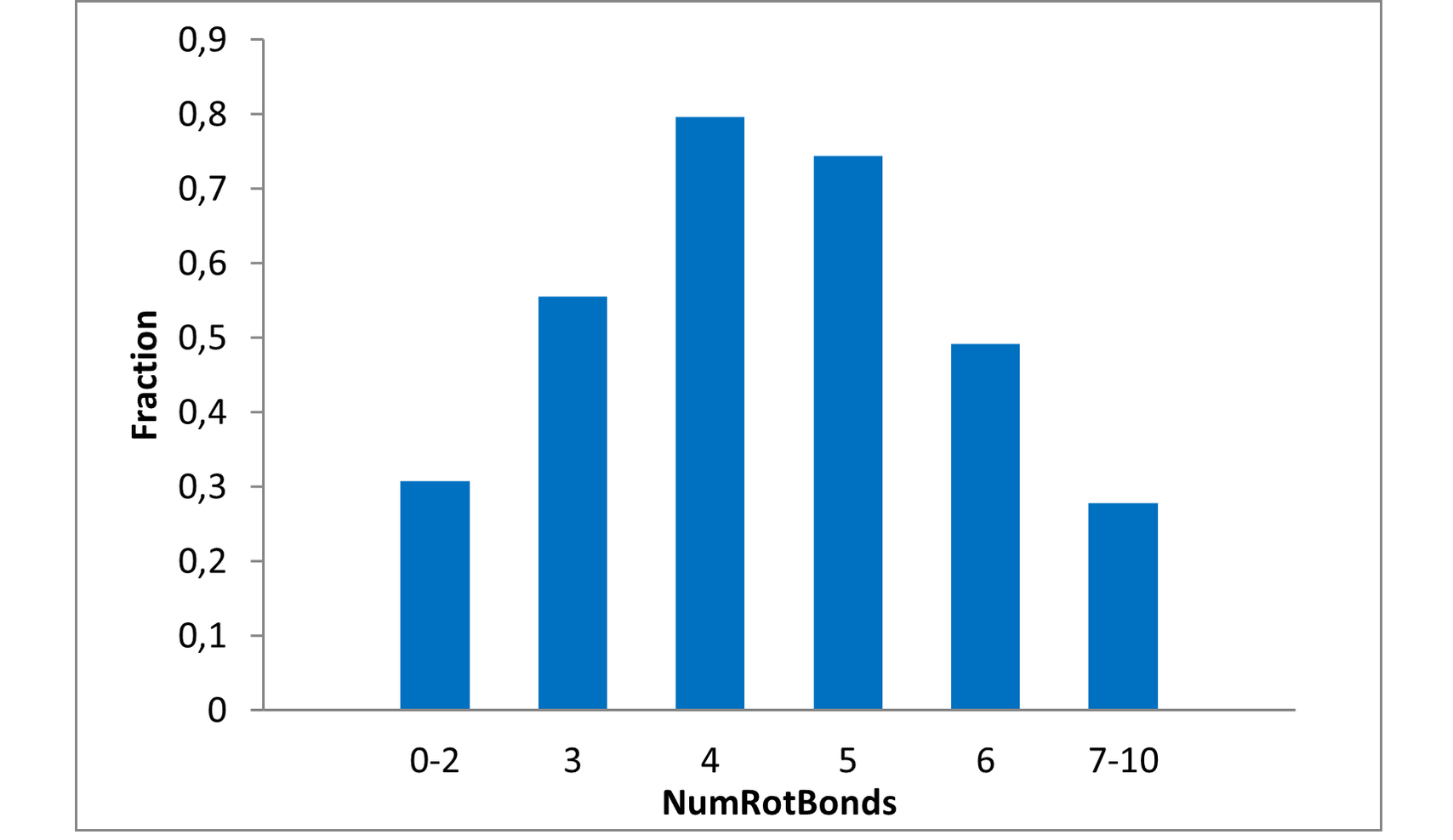
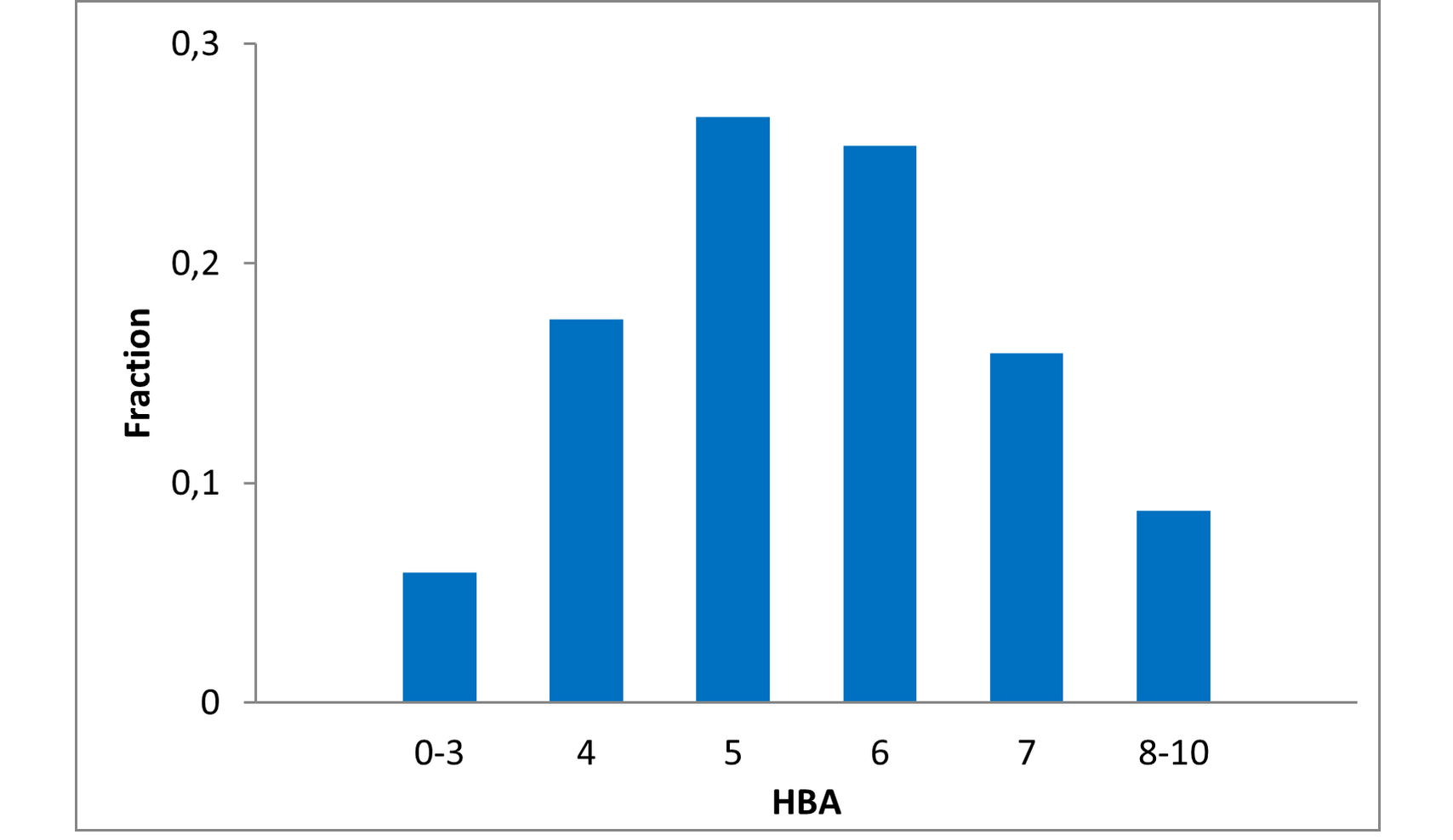
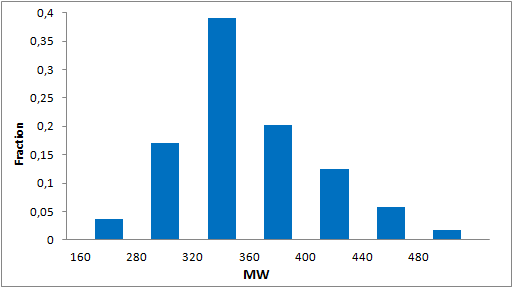
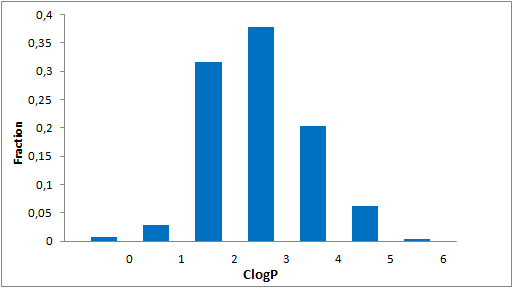
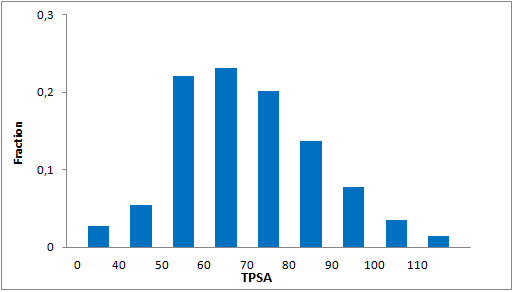
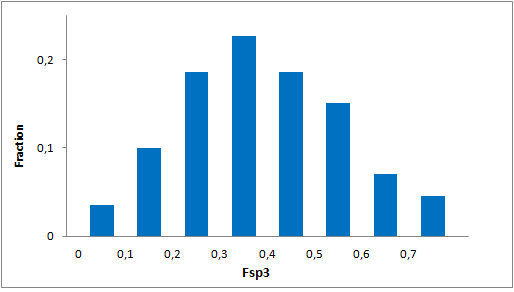
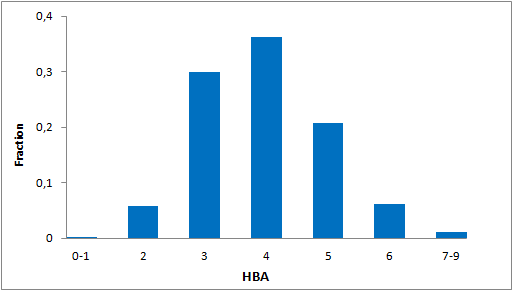
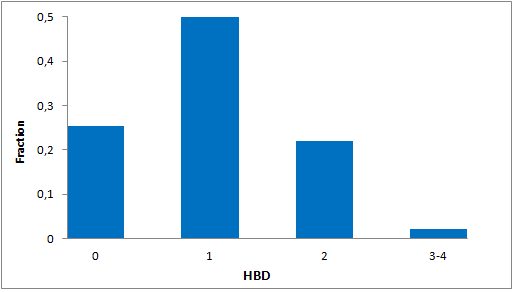
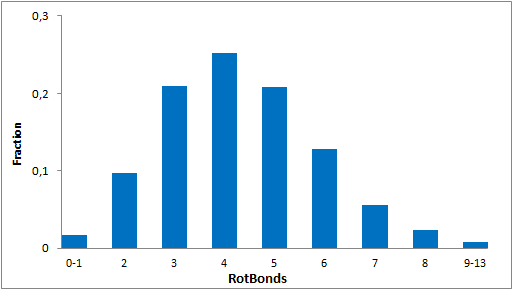
Structural parameters
We have identified several motifs/fragments/moieties which are abundant in agrochemicals. We used each of the following criteria for compound selection:
- contains 4 or more atoms of halogen (mostly Cl and F);
- has S-containing aliphatic/aromatic ring;
- contains polar, heteroatom-rich functional groups (e.g. diacyl hydrazines, N-acyl urea etc);
- contains oxazole/oxazoline rings;
- contains cyclopropanes;
- has O-containing aliphatic rings.
Structural exclusions
- No alcohols, primary amines and secondary amines occur less frequently in agrochemicals;
- No reactive/toxicophore groups (internal medchem filters). At the same time some compounds bearing active halogen (e.g. 2-chloropyridne) were left in the library;
- “Trivial” scaffolds and chemotypes.
Structural inclusions
- Acidic groups (e.g. carboxylic acids, acylsulfonamides etc.);
- Fsp3-rich molecules;
- Unusual/unique chemotypes/scaffolds/combination of cycles.
Diversity
The internal diversity filter was used to select the most diverse compounds (Coefficient of diversity: 0.86409).
Designed for discovery of new Nucleoside-like antivirals
3 200 compounds
In spite of significant success in medicine in last decades, development of effective new antiviral agents and vaccines continue to be a challenging task for the modern drug discovery. Viruses share most of the metabolic processes of host cells, thereby making difficult search of selective antiviral agents. However, some enzymes are only present in viruses and these are potential and most attractive targets for antiviral drugs.
For instance, there are several key enzymes, which are involved in the processes with nucleic acids like DNA- and RNA-polymerases. Also, reverse transcriptases possess high potential as antiviral targets. The success of previously approved anti-HCV drug Sofosbuvir has demonstrated the potency of small molecules and the important role of viral RNA-polymerases as drug targets. The series of new drug candidates with nucleoside-like scaffolds introduced by Gilead have shown promising results in treating of serious viral infections such as recently emerged Coronavirus, Ebola and RSV.
Typical Formats
Antiviral Library is available for supply in various pre-plated formats, including the following most popular ones:
Catalog No.
AVR-3200-0-Z-10
Compounds
3 200
3 plates
Amount
≤ 300 nL of 10 mM of DMSO solutions
Plates and formats
1536-well Echo LDV microplates, first and last four columns empty, 1280 compounds per plate
Price
Catalog No.
AVR-3200-10-Y-10
Compounds
3 200
10 plates
Amount
≤ 10 µL of 10 mM DMSO solutions
Plates and formats
384-well, Echo Qualified LDV microplates #001-12782 (LP-0200), first and last two columns empty, 320 compounds per plate
Price
Catalog No.
AVR-3200-50-Y-10
Compounds
3 200
10 plates
Amount
50 μL of 10 mM DMSO solutions
Plates and formats
384-well, Greiner Bio-One plates #781280, 1,2 and 23,24 columns empty, 320 compounds per plate
Price
Catalog No.
Library & follow-up package
Plates and formats
AVR-3200-10-Y-10 screening library 3 200 cmpds, hit resupply, analogs from 4.7M+ stock and synthesis from REAL Space
Price
*We will be happy to provide our library in any other most convenient for your project format. Please select among the following our standard microplates: Greiner Bio-One 781270, 784201, 781280, 651201 or Echo Qualified 001-12782 (LP-0200), 001-14555 (PP-0200), 001-6969 (LP-0400), C52621 or send your preferred labware. Compounds pooling can be provided upon request.
Download SD files
Examples of nucleoside mimetics as antiviral agents
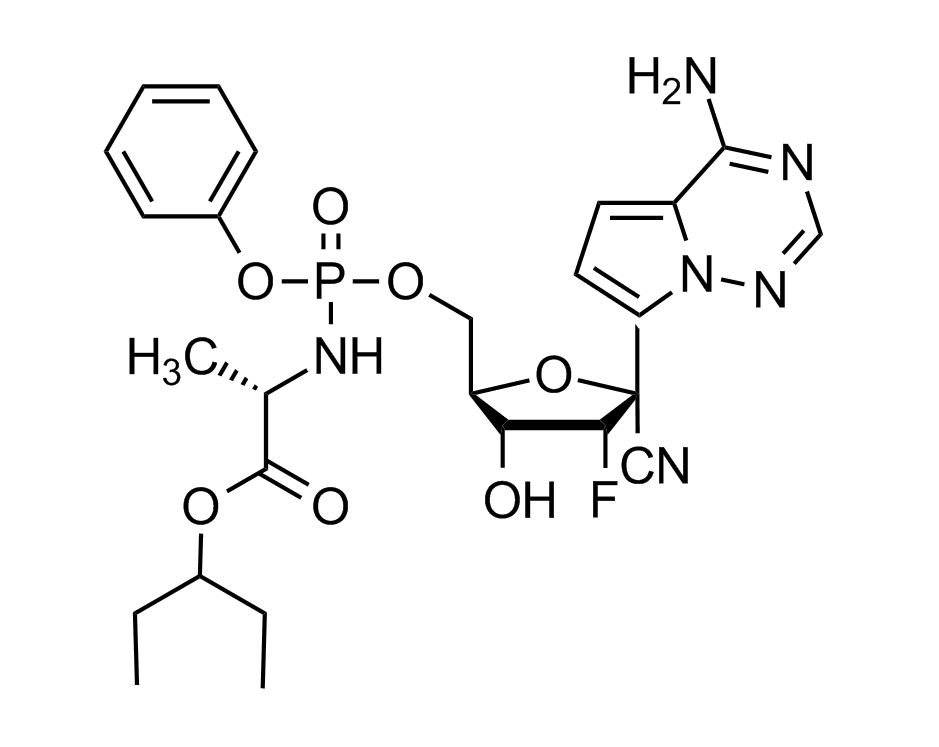
Remdesivir
Ebola, Junin and MERS viruses
studied against Coronaviruses, 2019-nCoV
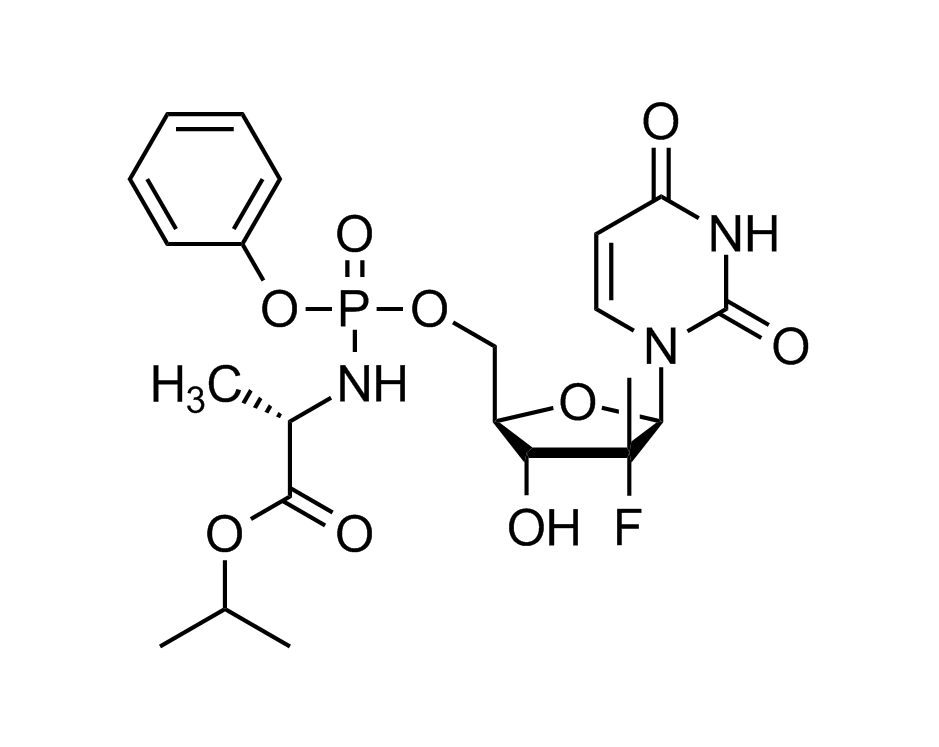
Sofosbuvir
Hepatitis C
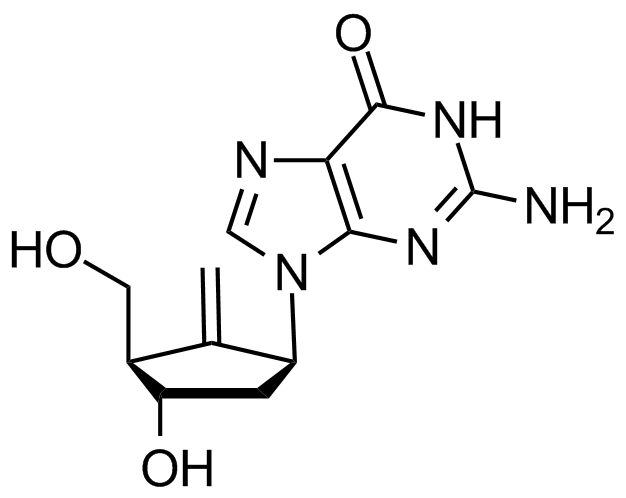
Entacavir
Hepatitis B virus
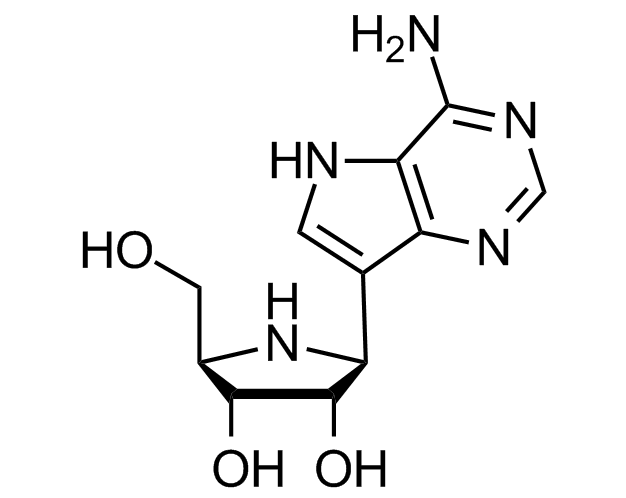
Galidesivir
Ebola & Marburg viruses
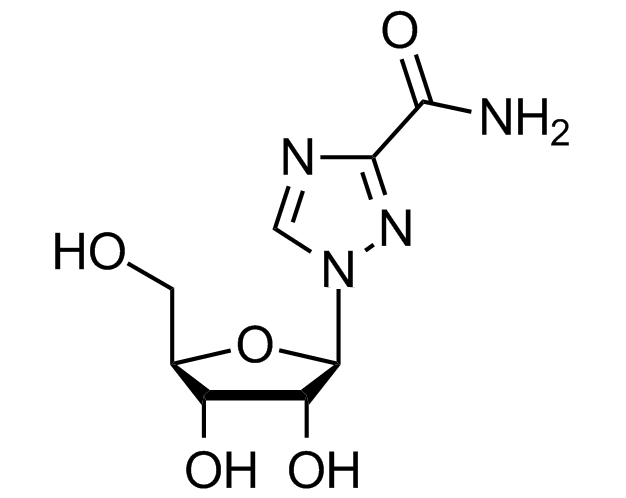
Ribavirin
RSV infection, hemorrhagic fevers
To meet growing needs in new antivirals Enamine has designed the library of nucleoside-like small molecules, enriched with recently synthesized series of nucleoside-like compounds. The library of carefully selected 3 200 compounds has been pre-plated for most convenient access and prompt delivery in various customized formats.
Library design
Analyzing the structures of known inhibitors we found that most of them are AVR-3200 side mimetics which are not distinguished from the nucleosides by virus enzymes in the active site.
Therefore taking into account pharmacophores and topology of nucleosides and reported nucleoside-like antiviral agents we carefully selected the set of Nucleoside mimetics from our screening collection. The compounds from the set contain natural-like moieties and diverse heterocycles as bioisosters of nucleosides. Additionally, special emphasis has been made on compounds that possess several H-bond donors and potentially can form similar interactions with the protein nucleoside-binding sites as a native nucleoside.
Examples of representative compounds from Antiviral Library
Designed for discovery of novel protein kinase inhibitors
64 960 compounds
Protein kinase inhibitors (PKI) represent an important and still emerging class of targeted therapeutic agents. It is difficult to overestimate the role of protein kinase inhibitors in modern anticancer and immune-oncology therapy. Almost 70 kinase inhibitors have received KNS-64960-4-Z-10 approval, with majority being approved for the treatment of different cancers, and about 200 kinase-targeted drugs are in clinical phase trials.
In order, to bring New Chemistry into this well-explored drug discovery field we carefully designed our Kinase targeted Library. The library is available in pre-plated format and can be promptly delivered in any custom formats. You can receive multiple benefits in the hit follow-up stages and lead optimization by using our Kinase Library:
- Hit confirmation support and hit expansion from dry stock of over 4.7M compounds.
- Straightforward and low-cost analogs synthesis through REAL Database technology.
- Hit-to-lead project support provided in timely manner with broad capabilities available on-site.
You have also an option to screen the librray directly at Enamine. We will be happy to offer you discount on the library cost depending on the collaboration scope.
Typical Formats
Kinase Library is available for supply in various pre-plated formats, including the following most popular ones:
Catalog No.
KNS-64-0-Z-10
Compounds
64 960
51 plates
Amount
≤ 300 nL of 10 mM of DMSO solutions
Plates and formats
1536-well Echo LDV microplates, first and last four columns empty, 1280 compounds per plate
Price
Catalog No.
KNS-64-10-Y-10
Compounds
64 960
203 plates
Amount
≤ 10 µL of 10 mM DMSO solutions
Plates and formats
384-well, Echo Qualified LDV microplates #001-12782 (LP-0200), first and last two columns empty, 320 compounds per plate
Price
Catalog No.
KNS-64-50-Y-10
Compounds
64 960
203 plates
Amount
50 μL of 10 mM DMSO solutions
Plates and formats
384-well, Greiner Bio-One plates #781280, first and last two columns empty, 320 compounds per plate
Price
Catalog No.
Library & follow-up package
Plates and formats
KNS-64-10-Y-10 screening library 64 960 cmpds, hit resupply, analogs from 4.7M+ stock and synthesis from REAL Space
Price
*We will be happy to provide our library in any other most convenient for your project format. Please select among the following our standard microplates: Greiner Bio-One 781270, 784201, 781280, 651201 or Echo Qualified 001-12782 (LP-0200), 001-14555 (PP-0200), 001-6969 (LP-0400), C52621 or send your preferred labware. Compounds pooling can be provided upon request.
Download SD file
Library design
In spite of extensive studies and impressive achievements in kinase inhibitor development there is a strong interest in development of novel and selective kinase inhibitors (including allosteric, covalent, bivalent) and subsequently qualitative kinases targeted libraries.
Our previous investigation and detailed structure analysis of known and most potent kinases’ inhibitors yielded the complex approach to design of unique kinase focused libraries. We used several approaches with proven record in development of known successful kinase inhibitors. Validated in silico screening along with selection of compounds bearing privileged scaffolds/moieties and bioisosteric core replacements were the key components of the dedicated design. We also introduced carefully selected compounds that were similar to known drugs.
Hinge Binders sublibrary - 24 000 compounds
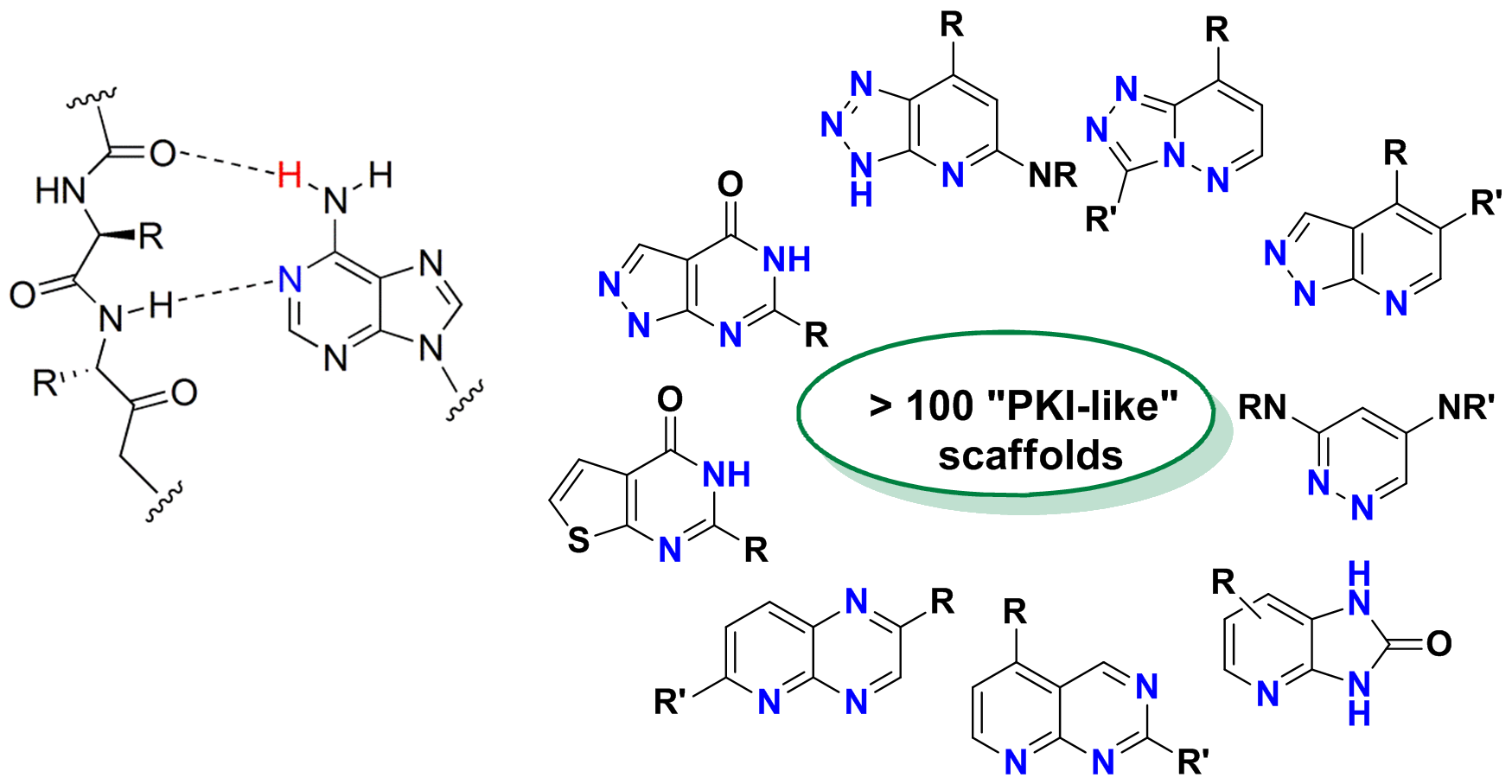
Well validated Cores with New Chemistry decoration
Pharmacophore & Shape similarity searches to the set of potent allosteric inhibitors
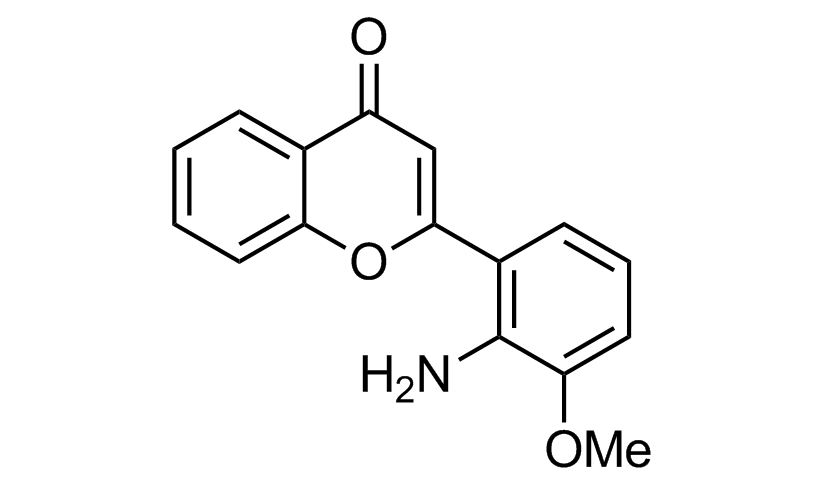
PD98059 (Allosteric inhibitor of MAPKK)
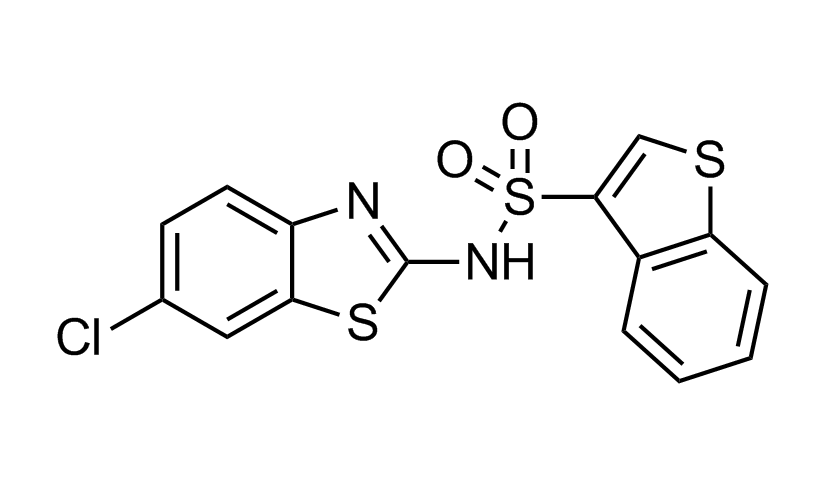
RS2 (allsoteric PDK1 inhibitor)
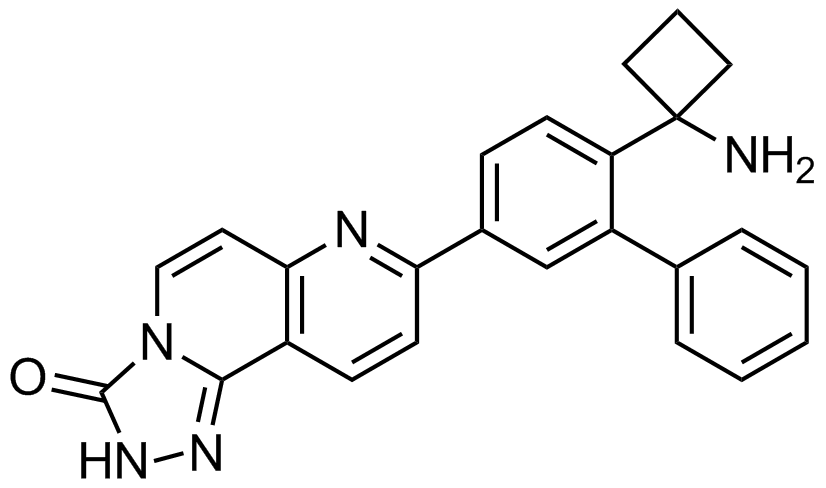
MK2206 (allosteric Akt-inhibitor)
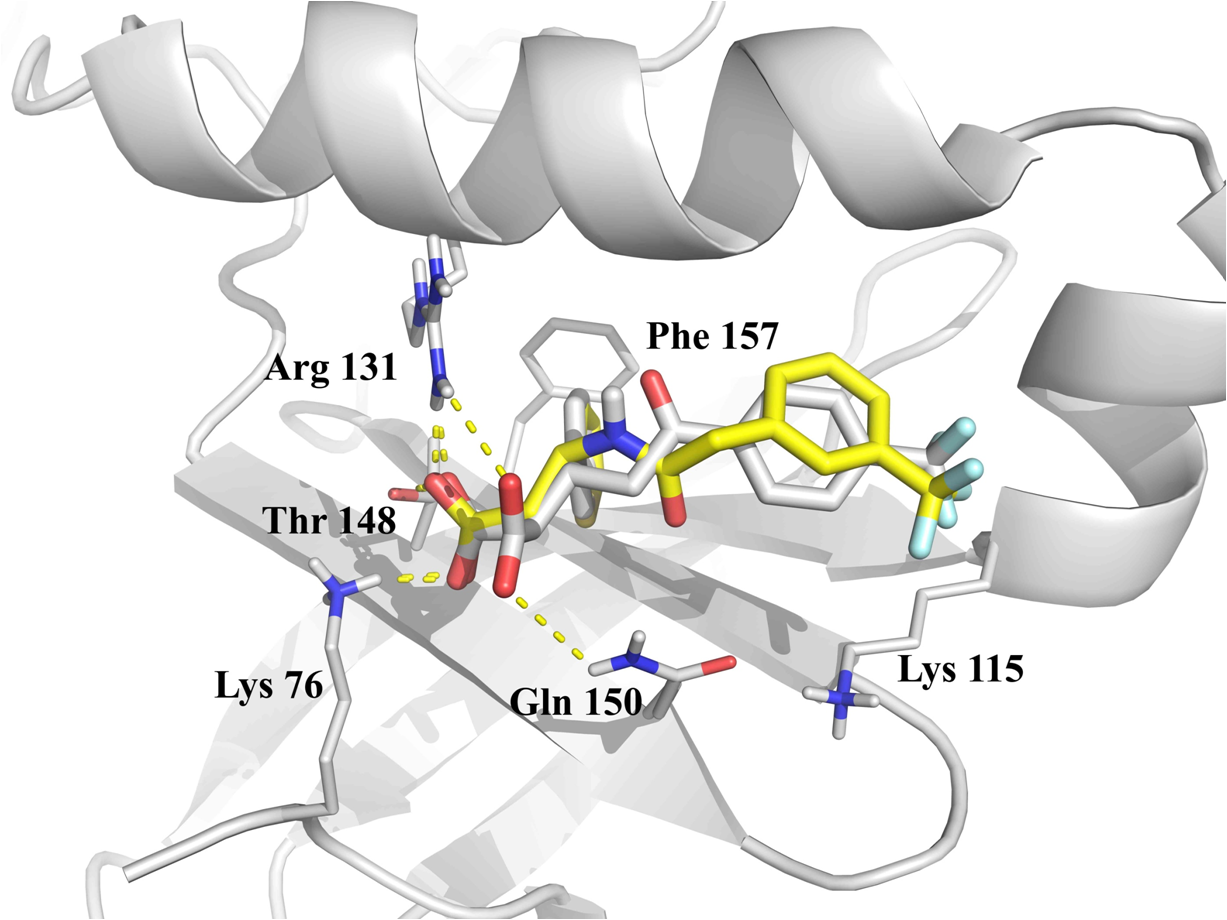
Docking into allosteric kinases binding sites
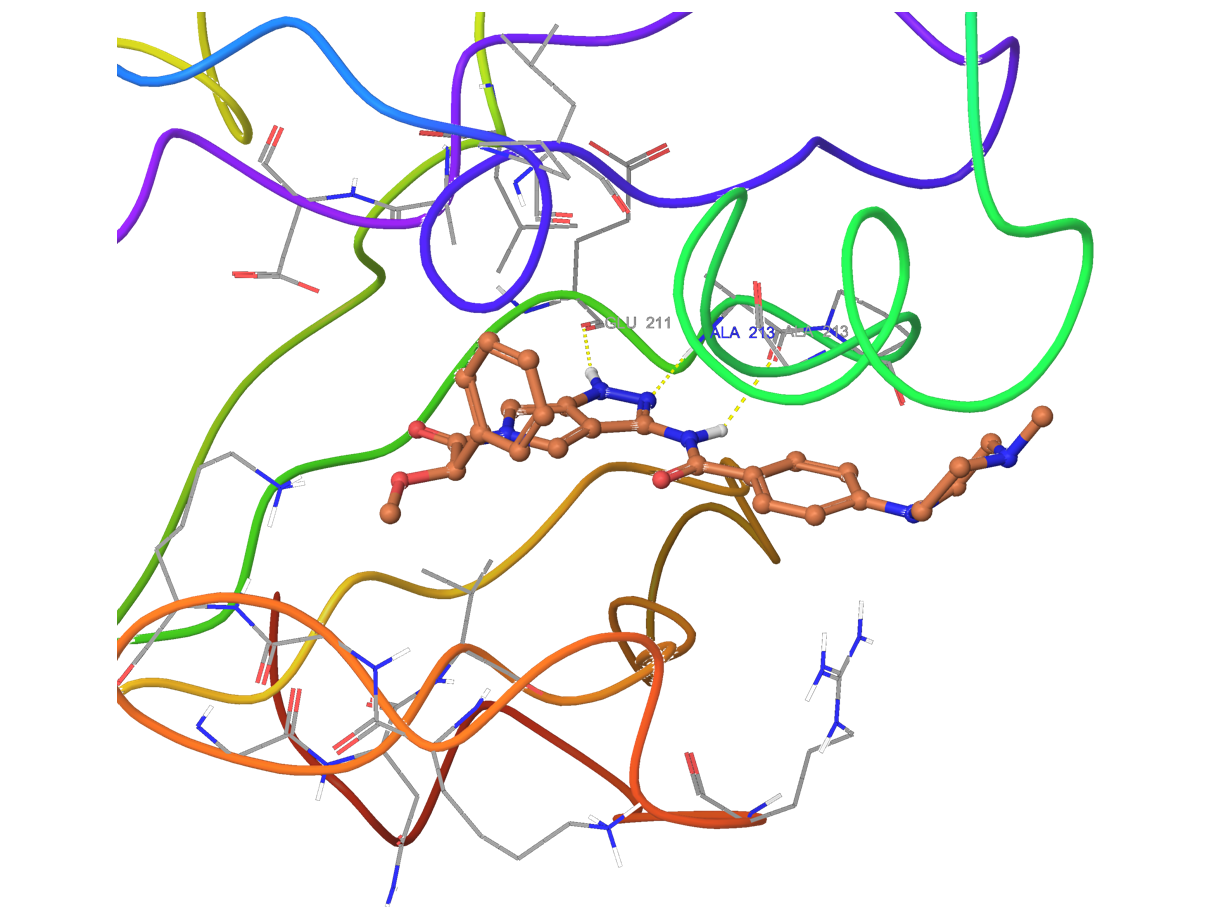
Docking calculations into ATP-binding pocket
Designed for discovery of novel PPI inhibitors
40 640 compounds
Having a pivotal role in multiple cellular processes, protein-protein interactions (PPIs) are responsible for the effects linked to the blockage of the substrate binding site. There are numerous examples of drug candidates that were withdrawn from clinical trials due to the unexpected effects of direct antagonists. This fact outlines the importance of developing new potent protein-protein interactions inhibitors.
We have carefully selected 40 640 diverse compounds specifically targeting PPIs. All compounds are stored as dry substances and they can be acquired in diverse custom formats. Using our PPI Library for hit discovery you receive multiple benefits allowing you to save on time and costs in lead generation:
- Dry stock of over 4.7M compounds for hit resupply and hit expansion.
- Low-cost synthesis of analogues within only 3 weeks through our REAL Database technology
- Medicinal chemistry support enhanced with on-site broad ADME/T panel
You have also an option to screen the Library directly at Enamine. In this case, we will be happy to offer discount on library cost depending on the collaboration scope.
Typical Formats
Protein-Protein Interaction Library is available for supply in various pre-plated formats, including the following most popular ones:
Catalog No.
PPI-40-0-Z-10
Compounds
40 640
32 plates
Amount
≤ 300 nL of 10 mM of DMSO solutions
Plates and formats
1536-well Echo LDV microplates, first and last four columns empty, 1280 compounds per plate
Price
Catalog No.
PPI-40-10-Y-10
Compounds
40 640
127 plates
Amount
10 µL of 10 mM DMSO solutions
Plates and formats
384-well, Echo Qualified LDV microplates #001-12782 (LP-0200), first and last two columns empty, 320 compounds per plate
Price
Catalog No.
PPI-40-50-Y-10
Compounds
40 640
127 plates
Amount
50 μL of 10 mM DMSO solutions
Plates and formats
384-well, Greiner Bio-One plates #781280, first and last two columns empty, 320 compounds per plate
Price
Catalog No.
Library & follow-up package
Plates and formats
PPI-40-10-Y-10 screening library 40 640 cmpds, hit resupply, analogs from 4.7M+ stock and synthesis from REAL Space
Price
*We will be happy to provide our library in any other most convenient for your project format. Please select among the following our standard microplates: Greiner Bio-One 781270, 784201, 781280, 651201 or Echo Qualified 001-12782 (LP-0200), 001-14555 (PP-0200), 001-6969 (LP-0400), C52621 or send your preferred labware. Compounds pooling can be provided upon request.
Download SD files
Library design
Systemic analysis of available structural data of numerous PPIs allowed us to develop dedicated approach to the library design. We have analyzed more than 20 different protein-protein complexes to highlight specific features of majority of potent inhibitors in this area. Several specific recognition patterns, like α-helix, β-sheet, PDZ-, PBD and bromodomains were used in the library design. As a result of Ligand- and Structure based in silico screening, we created the selection of compounds featuring:
- Specific recognition patterns, including hot spots analysis, key amino acids, secondary/tertiary structures, α-helices, ‘hot loops’ and specific protein domains affinity.
- Lead-like properties and sp3-rich core structural motifs. Compounds passed all including affiliated MedChem filters including PAINS.
- Latest chemistry and novel building blocks. Identified hits can be readily followed with synthesis of new analogs through REAL Database technology.
Examples of scaffolds populating PPI Library
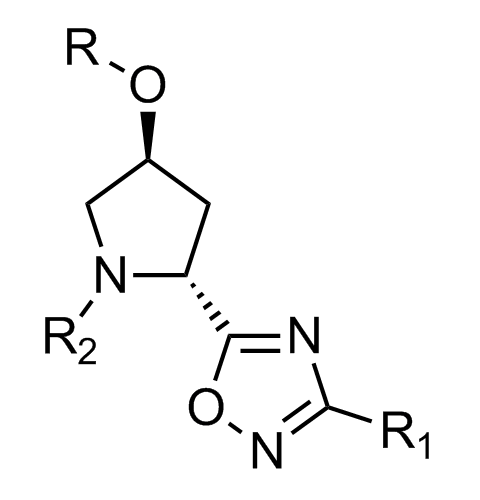
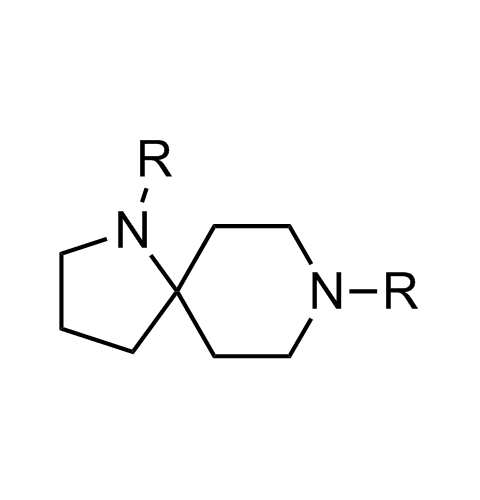
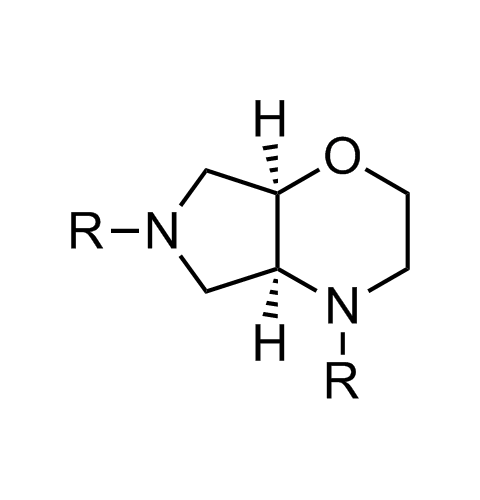
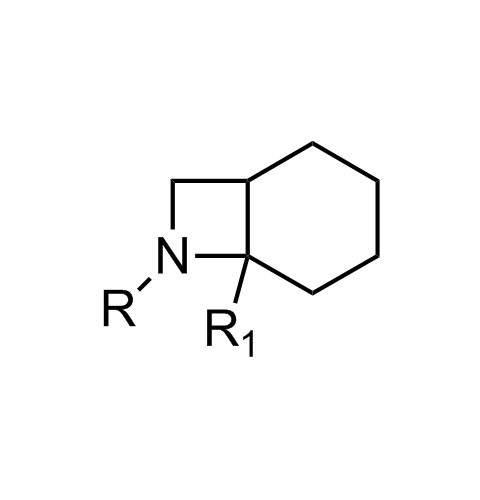
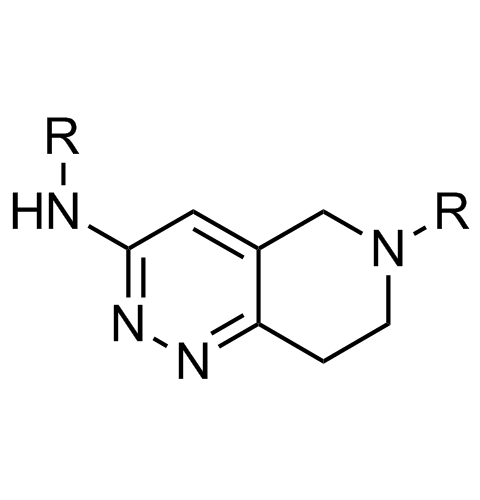
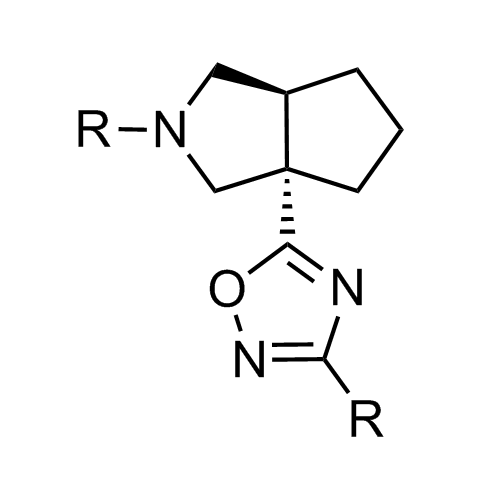
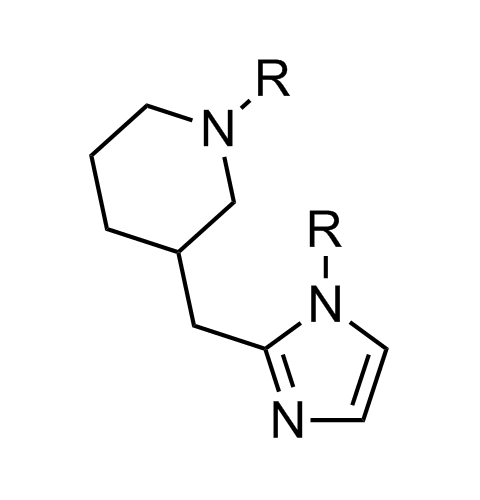
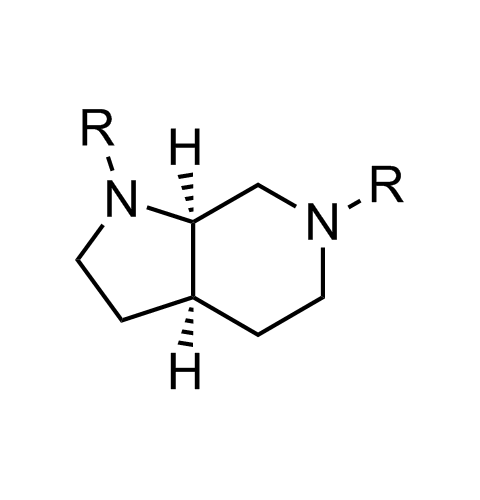
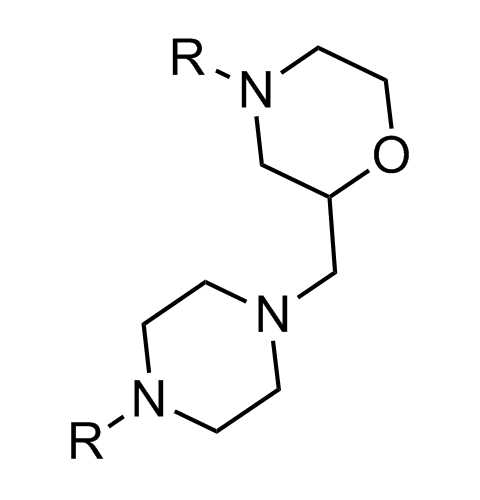
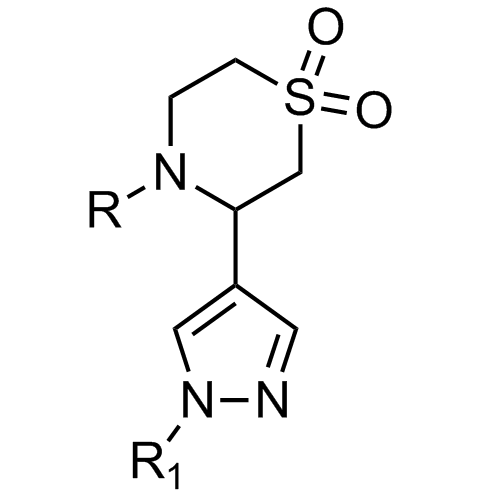
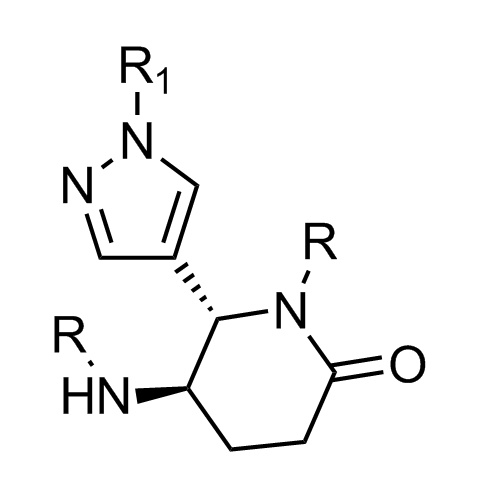
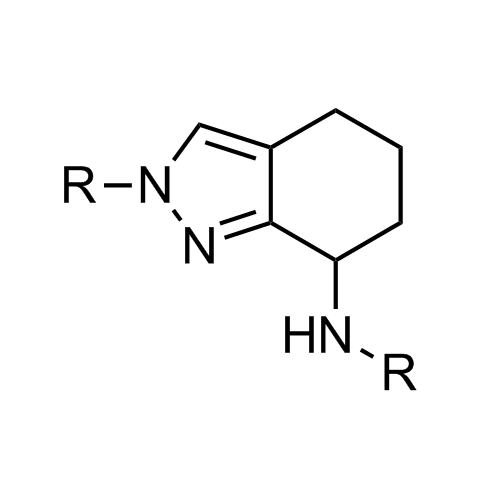
Importantly, our team of experienced computational, synthetic, medicinal chemists and biologists is ready to address your specific needs in tackling protein-protein interaction of your interest. Please, challenge us with your biological concepts, computational ideas and synthetic designs.
Library of compounds focusing to hit on a number of epigenetic targets
38 080 compounds
A solid molecular basis for the ways in which heritable information other than DNA sequence can regulate organism function is provided by recent findings in the area of epigenetics. Enamine is proud to provide our customers with compounds libraries focusing on several classes of epigenetic targets: Histone deacetylases (HDACs), Histone methyltransferases (HMT), DNA methyltransferase (DNMT), Bromodomain.
Download SD files
Epigenetics Library
Library code: EPG-38080
38 080 compounds
Version: 7 December 2023
Bromodomains
Library code: BRD-15
15 360 compounds
sublibrary of EPG-38080
Version: 7 December 2023
Histone Methyltransferase Library
10 560 compounds
sublibrary of EPG-38080
DNA Methyltransferase Library
6 880 compounds
sublibrary of EPG-38080
Histone Deacetylase focused Library
5 280 compounds
sublibrary of EPG-38080
Library design
Due to the extremely wide variety among epigenetic target classes, different approaches were used to create the library. Each class of epigenetic targets is represented with diverse families, which should be treated individually. When creating the libraries main idea was to make family-specific compound libraries, rather than particular molecular target oriented. Such an approach is based on dividing molecular targets from similar family into bunch of clusters according to the information of their binding sites, including site’s spatial structure, amino-acid composition etc., thus allowing us to build a profile describing the family of targets and specificity of their interactions with ligands. This data provides a strong basis for selecting both: the most representative, centroid protein structure and allows building preliminary pharmacophore model to filter out those compounds not having sufficient structural features to show good binding. The next step includes advanced 3D pharmacophore model creation in order to further decrease the number of compounds to be subjected to docking procedure. Next comes docking with the final filtering and inspection of obtained results.
The library for each target class was created utilizing combined method, which includes both ligand-based as well as receptor-based approaches. Thus, generic flow chart may be represented as follows:
- Analysis of all available structural data (PDB, literature sources etc.)
- Pharcophore models creating and validation
- Advanced pharmacophore search, utilizing key points of interactions, restricted volumes and spatial conformation requirements
- Molecular docking
- Precise tuning of obtained results, visual inspection
Such a method allows us to create libraries which may come as invaluable tool for early stages study of poorly explored targets, giving an opportunity to reveal compounds with high affinity to the target. Therefore providing solid basis for the further hit exploration step, during which the researcher may focus on increasing the specificity of obtained hit compounds towards other representatives of the targeted family.
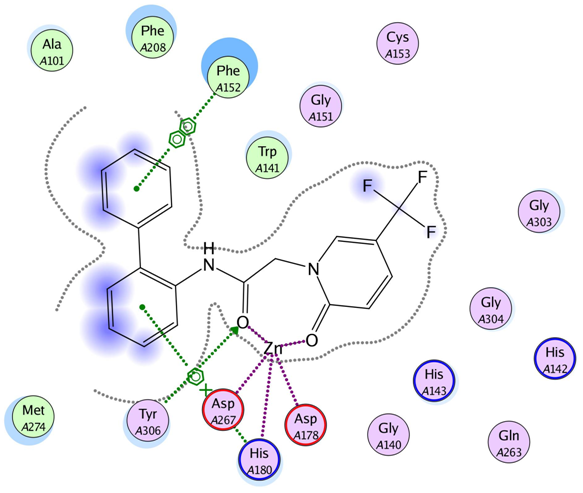
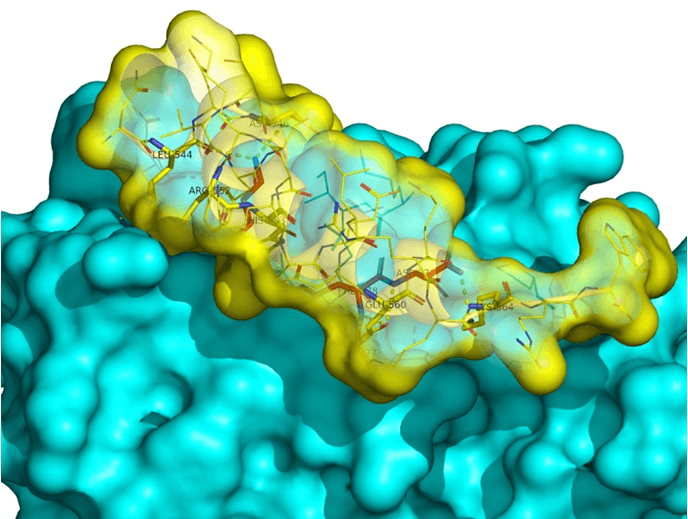
HDACs pattern of the ligand-receptor interaction.