Special selection of Serine focused irreversible binders
1 600 compounds
The Serine hydrolase enzyme family is one of the largest and most diverse protein classes, including proteases, lipases, esterases, thioesterases, amidases and peptidases. All these enzymes utilize a base-activated Serine residue cleavage of amide bonds in substrates via a covalent acyl-enzyme intermediate.
Different specific covalent-acting chemical probes have increasingly been used in proteome-wide target identification and imaging and for finding inhibitors with high specificity among related enzymes and enzyme isoforms. Significant number of known covalent drugs and natural products have approved efficacy of such approach in drug discovery.
Our library of Serine focused covalent fragments is available in versatile pre-plated formats for most convenient and fast delivery. All compounds passed QC check (90%+ purity) before formatting to ensure quality of the library.
Typical Formats
Serine-Focused Covalent Library is available for supply in various pre-plated formats, including the following most popular ones:
Catalog No.
SER-1-0-Z-10
Compounds
1 600
2 plates
Amount
≤ 300 nL of 10 mM of DMSO solutions
Plates and formats
1536-well Echo LDV microplates, first and last four columns empty, 1280 compounds per plate
Price
Catalog No.
SER-1-10-Y-10
Compounds
1 600
5 plates
Amount
10 µL of 10 mM DMSO solutions
Plates and formats
384-well, Echo Qualified LDV microplates #001-12782 (LP-0200), first and last two columns empty, 320 compounds per plate
Price
Catalog No.
SER-1-50-Y-10
Compounds
1 600
5 plates
Amount
50 μL of 10 mM DMSO solutions
Plates and formats
384-well, Greiner Bio-One plates #781280, first and last two columns empty, 320 compounds per plate
Price
Catalog No.
Library & follow-up package
Plates and formats
SER-1-10-Y-10 screening library 1 600 cmpds, hit resupply, analogs from 4.7M+ stock and synthesis from REAL Space
Price
*We will be happy to provide our library in any other most convenient for your project format. Please select among the following our standard microplates: Greiner Bio-One 781270, 784201, 781280, 651201 or Echo Qualified 001-12782 (LP-0200), 001-14555 (PP-0200), 001-6969 (LP-0400), C52621 or send your preferred labware. Compounds pooling can be provided upon request.
Download SD file
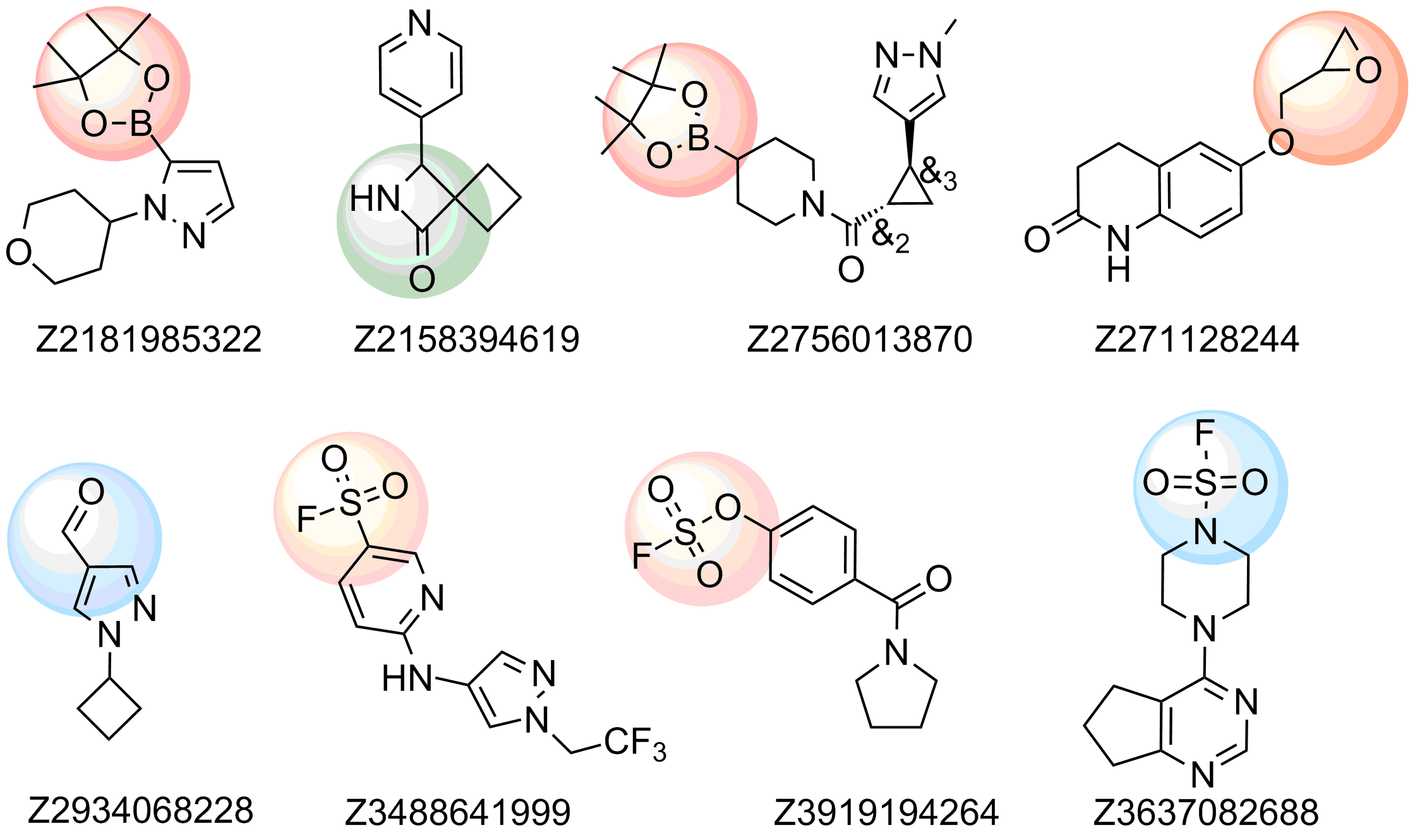
Examples of compounds in the library
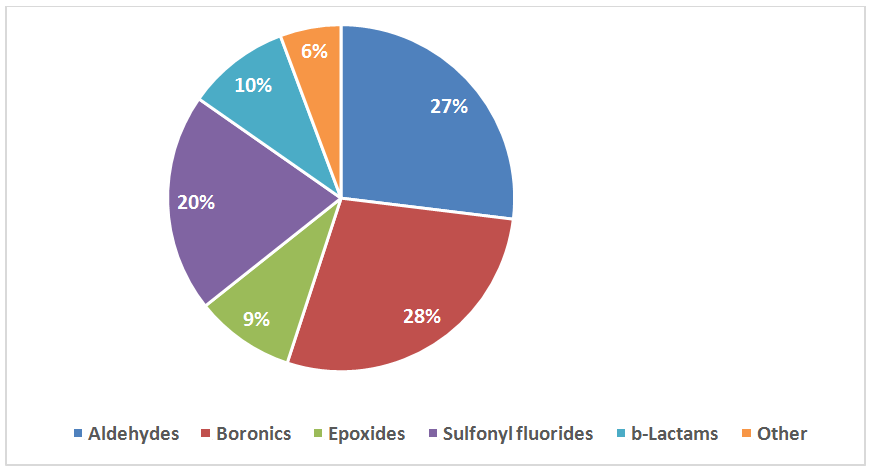
Library Design
Recently elaborated approaches to parallel synthesis, allowed Enamine to synthesize the largest commercial source of covalent binders. We focused our efforts on synthesis of compounds with most reliable covalent warheads, with well determined reactivity. Our Serine focused library was designed based on a combination of specific moieties, reported to form covalent bonds particularly with Serine residue and presence of a drug-like, MedChem friendly core in a molecule. The library has also been shaped with Ro3 criteria to meet all requirements of FBDD.
The following functional groups, reported to form covalent bonds in proteins with Ser residue, have been used for construction of the library:
- Sulfonyl fluorides, fluorosulfonates and sulfamoyl fluorides
- Epoxides
- β-lactams and β-Propiolactone
- Boronic acids and pinacolates
- Aldehydes
Hits derived from this library can be easily followed with analogues from over 4.7M stock compounds or either synthesis of new compounds through REAL Database technology within 3 weeks only.
We provide Hit Confirmation support for all our libraries with the samples resupply from dry powders, QC check and HPLC repurification. Hits can be resynthesized in 2 weeks only.



